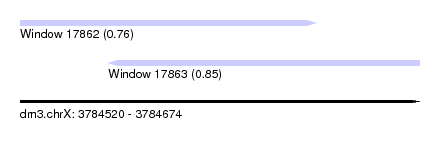

| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,784,520 – 3,784,674 |
| Length | 154 |
| Max. P | 0.854890 |

| Location | 3,784,520 – 3,784,634 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.61 |
| Shannon entropy | 0.18155 |
| G+C content | 0.46349 |
| Mean single sequence MFE | -36.11 |
| Consensus MFE | -26.33 |
| Energy contribution | -28.45 |
| Covariance contribution | 2.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.756319 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 3784520 114 + 22422827 -----GUGAAGUGACCC-ACUCGGUUGCUGGACUUCAUUUACAUCUAAAUCGGAAUGCUGGCCUUUUUCGGGCUUUUGCGGCCAACAUUUCUGAUUUGAUUGAGGUUUUGCAGUCAGUGC -----...........(-(((.(..(((..(((((((.....(((.(((((((((((.(((((........((....))))))).)).)))))))))))))))))))..)))..))))). ( -37.90, z-score = -2.59, R) >droEre2.scaffold_4690 1168256 119 + 18748788 UUUCUGCGAAAUGCCCCCACUACCUUCCUGGACUUAAUUUACAUCUAAAUCGGAAUGUUGGCCUUUAUCGCGCUUUUGCGGCCAACAUUUCUGAUUUUAUUGAGGCUU-GCAGGCUCUGC ..(((((((...(((.......((.....))..........((...(((((((((((((((((........((....)))))))))).)))))))))...)).)))))-)))))...... ( -36.20, z-score = -3.61, R) >droYak2.chrX 17140526 118 - 21770863 UUUCUGCGAAGUGACCC-ACUACGUUCCUGGACUUCAUUUACAUCUAAAUCGGAGUGCUGGCAUUUUUCGGGAUUUUGCGGCCAACAUUUCUGAUUUGAUUGAGGUUU-GCAGUCUCUGC ...(((((.((((...)-))).)......((((((((.....(((.(((((((((((.((((..................)))).)).))))))))))))))))))))-))))....... ( -28.87, z-score = -0.18, R) >droSec1.super_4 3255922 119 - 6179234 UCUCUGCGAAGUGACCC-ACUCGGUUGCUGGACUUCAUUUACAUCUAAAUCGGAAUGCUGGCCUUUUUCGGGCUUUUGCGGCCAACAUUUCUGAUUUGAUUGGGGUUUUGCAGUCAGUGC ...((((((((..(((.-....)))..)).(((((((.....(((.(((((((((((.(((((........((....))))))).)).)))))))))))))))))))))))))....... ( -38.30, z-score = -2.06, R) >droSim1.chrX 2791218 119 + 17042790 UCUCUGCGAAGUGACCC-ACUCGGUUGCUGGACUUCAUUUACAUCUAAAUCGGAAUGCUGGCCUUUUUCGGGCUUUUGCGGCCAACAUUUCUGAUUUGAUUGAGGUUUUGCAGUCAGUGC ...((((((((..(((.-....)))..)).(((((((.....(((.(((((((((((.(((((........((....))))))).)).)))))))))))))))))))))))))....... ( -39.30, z-score = -2.56, R) >consensus UCUCUGCGAAGUGACCC_ACUCGGUUGCUGGACUUCAUUUACAUCUAAAUCGGAAUGCUGGCCUUUUUCGGGCUUUUGCGGCCAACAUUUCUGAUUUGAUUGAGGUUUUGCAGUCAGUGC ...((((..(((((((......)))))))((((((((.....(((.(((((((((((.(((((................))))).)).)))))))))))))))))))).))))....... (-26.33 = -28.45 + 2.12)

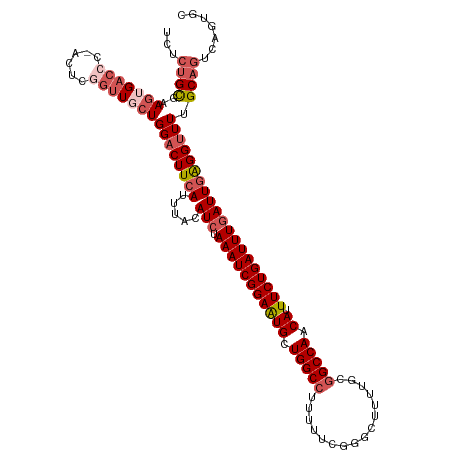

| Location | 3,784,554 – 3,784,674 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.33 |
| Shannon entropy | 0.17346 |
| G+C content | 0.48905 |
| Mean single sequence MFE | -36.17 |
| Consensus MFE | -28.15 |
| Energy contribution | -29.27 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.854890 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 3784554 120 - 22422827 AAGGCCCUGGAACGGGAAACGUGCCACAGGUGCGAAAACAGCACUGACUGCAAAACCUCAAUCAAAUCAGAAAUGUUGGCCGCAAAAGCCCGAAAAAGGCCAGCAUUCCGAUUUAGAUGU .((((((......))).....(((....(((((.......)))))....)))...)))..((((((((.(((.((((((((................))))))))))).))))).))).. ( -35.99, z-score = -1.83, R) >droEre2.scaffold_4690 1168296 118 - 18748788 GUGGGCUUUGAACG-GAAAGGUGCCAUAGGAGCGAAAGCAGCAGAGCCUGC-AAGCCUCAAUAAAAUCAGAAAUGUUGGCCGCAAAAGCGCGAUAAAGGCCAACAUUCCGAUUUAGAUGU (..(((((((...(-(.......))......((....))..)))))))..)-...........(((((.(((.((((((((................))))))))))).)))))...... ( -39.69, z-score = -2.68, R) >droYak2.chrX 17140565 119 + 21770863 GAGGACCUGGAAUGCAAAACGUGCUACAGGAGCGAAAACAGCAGAGACUGC-AAACCUCAAUCAAAUCAGAAAUGUUGGCCGCAAAAUCCCGAAAAAUGCCAGCACUCCGAUUUAGAUGU ((((.((((....(((.....)))..))))..........((((...))))-...)))).((((((((.((..(((((((..................))))))).)).))))).))).. ( -31.87, z-score = -2.42, R) >droSec1.super_4 3255961 120 + 6179234 GAGGGCCUGGAACGGGAAACGUGCCACAGGAGCGAAAACAGCACUGACUGCAAAACCCCAAUCAAAUCAGAAAUGUUGGCCGCAAAAGCCCGAAAAAGGCCAGCAUUCCGAUUUAGAUGU ..(((((((..(((.....)))....))))..........(((.....))).....))).((((((((.(((.((((((((................))))))))))).))))).))).. ( -35.59, z-score = -1.74, R) >droSim1.chrX 2791257 120 - 17042790 GAGGGCCUGGAACGGGAAACGUGCCACAGGAGCGAAAACAGCACUGACUGCAAAACCUCAAUCAAAUCAGAAAUGUUGGCCGCAAAAGCCCGAAAAAGGCCAGCAUUCCGAUUUAGAUGU ((((.((((..(((.....)))....))))..........(((.....)))....)))).((((((((.(((.((((((((................))))))))))).))))).))).. ( -37.69, z-score = -2.35, R) >consensus GAGGGCCUGGAACGGGAAACGUGCCACAGGAGCGAAAACAGCACUGACUGCAAAACCUCAAUCAAAUCAGAAAUGUUGGCCGCAAAAGCCCGAAAAAGGCCAGCAUUCCGAUUUAGAUGU ((((.((((..(((.....)))....))))..........(((.....)))....)))).((((((((.(((.((((((((................))))))))))).))))).))).. (-28.15 = -29.27 + 1.12)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:12:40 2011