| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,727,778 – 3,727,876 |
| Length | 98 |
| Max. P | 0.930079 |

| Location | 3,727,778 – 3,727,876 |
|---|---|
| Length | 98 |
| Sequences | 7 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 56.89 |
| Shannon entropy | 0.77084 |
| G+C content | 0.38734 |
| Mean single sequence MFE | -16.93 |
| Consensus MFE | -6.25 |
| Energy contribution | -6.41 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.35 |
| SVM RNA-class probability | 0.930079 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
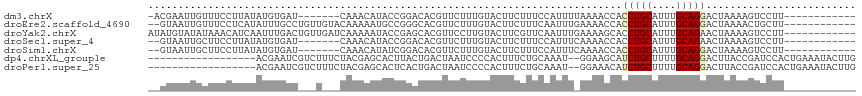
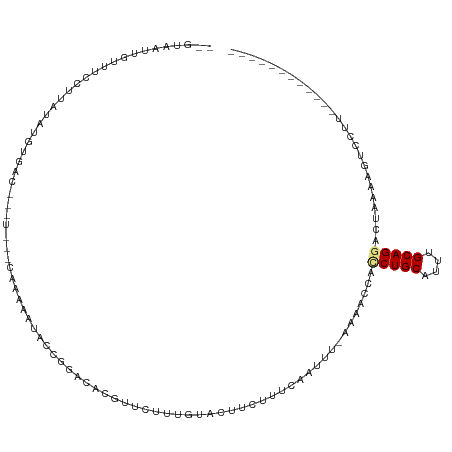
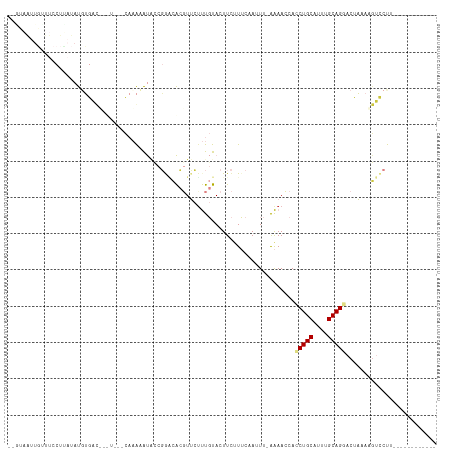
>dm3.chrX 3727778 98 - 22422827 -ACGAAUUGUUUCCUUAUAUGUGAU-------CAAACAUACCGGACACGUUCUUUGUACUUCUUUCCAUUUUAAAACCACCUGCAUUUGCAGGACUAAAAGUCCUU------------ -(((((.(((.(((...(((((...-------...)))))..))).)))...)))))......................(((((....))))).............------------ ( -18.60, z-score = -2.18, R) >droEre2.scaffold_4690 1111128 104 - 18748788 --GUAAUUGUUUCCUCAUAUUUGCCUGUUGUACAAAAAUGCCGGGCACGUUCUUUGUACUUCUUUCAAUUUGAAAACCACCUGCAUUUGCAGGACUAAAACUGCUU------------ --(((...((((..(((...(((...(..(((((((((((.......)))).)))))))..)...)))..)))))))..(((((....)))))........)))..------------ ( -19.10, z-score = -0.70, R) >droYak2.chrX 17081249 106 + 21770863 AUAUGUAUAUAAACAUCAAUUUGACUGUUGAUCAAAAAUACCGAGCACGUUCCUUGUACUUCGUUCAAUUUGAAAAGCACCUGCAUUUGCAGAACUAAAAGUCCUU------------ ..((((......))))......(((((((..(((((...((.(((.(((.....))).))).))....)))))..)))..((((....)))).......))))...------------ ( -17.40, z-score = -1.02, R) >droSec1.super_4 3193964 97 + 6179234 --GUAAUUGCUUCCUUAUAUGUGAU-------CAAACAUACCGGACACGUUCUUUGUACUUCUUUCCAUUUCAAAACCACCUGCAUUUGCAGAACUAAAAGUCCUU------------ --...............(((((...-------...)))))..(((((((.....))).......................((((....))))........))))..------------ ( -14.50, z-score = -1.59, R) >droSim1.chrX 2733583 97 - 17042790 --GUAAUUGCUUCCUUAUAUGUGAU-------CAAACAUAUCGGACACGUUCUUUGUACUUCUUUCCAUUUCAAAACCACCUGCAUUUGCAGGACUAAAAGUCCUU------------ --((((.(((.(((..((((((...-------...)))))).))).(((.....))).........................))).))))(((((.....))))).------------ ( -19.30, z-score = -2.76, R) >dp4.chrXL_group1e 6866301 98 - 12523060 ------------------ACGAAUCGUCUUUCUACGAGCACUUACUGACUAAUCCCCACUUUCUGCAAAU--GGAAGCAUCUGCUUUUGCAGGACUUACCGAUCCACUGAAAUACUUG ------------------.((..((((......)))).......................(((((((...--.(((((....)))))))))))).....))................. ( -15.00, z-score = -0.74, R) >droPer1.super_25 201763 98 - 1448063 ------------------ACGAAUCGUCUUUCUACGAGCACUCACUGACUAAUCCCCACUUUCUGCAAAU--GGAAACAUCUGCUUUUGCAGGACUUACCGAUCCACUGAAAUACUUG ------------------.....((((......))))....(((.((....(((......((((((((((--(....)))......))))))))......))).)).)))........ ( -14.60, z-score = -1.12, R) >consensus __GUAAUUGUUUCCUUAUAUGUGAC___U___CAAAAAUACCGGACACGUUCUUUGUACUUCUUUCAAUUU_AAAACCACCUGCAUUUGCAGGACUAAAAGUCCUU____________ ...............................................................................(((((....)))))......................... ( -6.25 = -6.41 + 0.16)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:12:22 2011