| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,700,373 – 3,700,538 |
| Length | 165 |
| Max. P | 0.993120 |

| Location | 3,700,373 – 3,700,538 |
|---|---|
| Length | 165 |
| Sequences | 7 |
| Columns | 172 |
| Reading direction | forward |
| Mean pairwise identity | 69.53 |
| Shannon entropy | 0.56186 |
| G+C content | 0.47542 |
| Mean single sequence MFE | -53.94 |
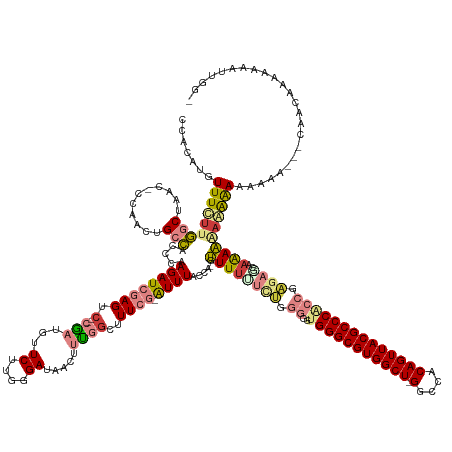
| Consensus MFE | -21.87 |
| Energy contribution | -23.59 |
| Covariance contribution | 1.72 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.75 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.59 |
| SVM RNA-class probability | 0.993120 |
| Prediction | RNA |
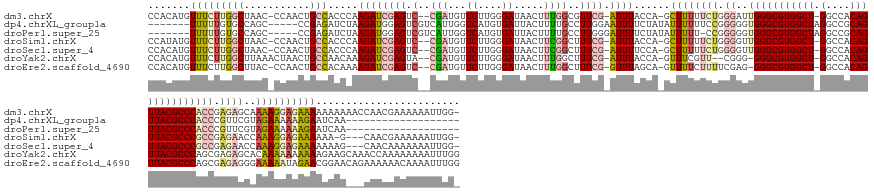
Download alignment: ClustalW | MAF
>dm3.chrX 3700373 165 + 22422827 CCACAUGUUUCUUGGCUAAC-CCAACUGCCACCCAAGAUCGAGUC--CGAUGUUCUUGGGAUAACUUUGGCGUUCG-AUUUACCA-GCUUUUUCUGGGAUUGGGCGUGGCU-GGCCACAGUUACGCCCACCGAGAGCAAAAGGAGAAAAAAAAAACCAACGAAAAAAUUGG- .......(((((((((....-......))).((((((((((....--)))...)))))))....((((.((.((((-.....(((-(......))))...(((((((((((-(....)))))))))))).)))).)).)))))))))).......((((........))))- ( -56.50, z-score = -3.07, R) >dp4.chrXL_group1a 6362917 141 + 9151740 -------UUUUUGUGCCAGC-----CCGAGAUCUAAGAUGGAGUCGUCAUUGGUCAUGUGAUUACUUUUGCCUUGGAAUUUUCUAUAUUUUUUCCGGGGGUGGGCGUGGCUAGGCCGCAGUUACGCCCACCCGUUCGUAGAAAAAAGAAUCAA------------------- -------(((((.(((.(((-----(((.((((((((..(((((.(((((.......))))).)))))...))))))...............)))))((((((((((((((.......))))))))))))))))).))).)))))........------------------- ( -49.66, z-score = -2.76, R) >droPer1.super_25 487310 140 - 1448063 -------UUUUUGUGCCAGC-----CCGAGAUCUAAGAUGGAGUCGUCAUUGGUCAUGUGAUUACUUUUGCCUUGGGAUUUUCUAUAUUUUU-CCGGGGGUGGGCGUGGCUAGGCCGCAGUUACGCCCACCCGUUCGUAGAAAAAAGAAUCAA------------------- -------(((((.(((.(((-----(((.((((((((..(((((.(((((.......))))).)))))...))))))..............)-))))((((((((((((((.......))))))))))))))))).))).)))))........------------------- ( -48.83, z-score = -2.25, R) >droSim1.chrX 2713150 161 + 17042790 CCAUAUGUUUCUUGGCUAAC-CCAACUGCCACCCAAGAUCGAGUC--CGAUGUUCUUGGGAUAACUUCGGCUUUCG-AUUUACCA-GCUUUUUCUGGGGUUGGGCGUGGCU-GGCCACAGUUACGCCCGCCGAGAACCAAAGGAGAAAAAA-G---CAACGAAAAAAUUGG- (((....((((.((((....-......))))((((((((((....--)))...)))))))....(((.((.((((.-.....(((-(......))))(((.((((((((((-(....)))))))))))))))))).)).))).........-.---....))))....)))- ( -57.60, z-score = -2.94, R) >droSec1.super_4 3167420 162 - 6179234 CCACAUGUUUCUUGGCUAAC-CCAACUGCCACCCAAGAUCGAGUC--CGAUGUUCUUGGGAUAACUUCGGCUUUCG-AUUUUCCA-GCUUUUUCUGGGGUUGGGCGUGGCU-GGCCACAGUUACGCCCGCCGAGAACCAAAGGAGAAAAAAAG---CAACAAAAAAAUUGG- ((...((.((((((((.(((-((...........(((((((((.(--(((.(((........))).))))..))))-))))).((-(......))))))))((((((((((-(....))))))))))))))))))).))..))..........---...............- ( -57.50, z-score = -2.77, R) >droYak2.chrX 17052668 164 - 21770863 CCACAUGUUUCUUGGCUUAAACUAACUGCCAACAAAGAUCGAGUA--CGAUGUUCUUGGGAUAACUUUGGCUUUCG-AUUUACCA-GUUUCGUU--CGGG-GGGCGUGGCU-GGCCACAGUUACGCCCAGCGAGAGCACAAAAAAAAAAGAAGCAAACCAAAAAAAAUUUGG .....(((((((((((...........)))..(((((((((....--))))(((.....)))..)))))(((((((-.....((.-(.......--).))-((((((((((-(....)))))))))))..)))))))..........))))))))..(((((.....))))) ( -52.00, z-score = -2.81, R) >droEre2.scaffold_4690 1083714 165 + 18748788 CCACAUGUUUCUUGGCUUAC-CCAACUGCCACAAAAGAUCGAGUC--CGAUGUUCUUGGGAUAACUUUGGCUUUCG-GUUUAGCA-GUUUUCUUUUCGAG-GGGCGUGGCU-GGCCACAGUUACGCCCAGCGAGAGGGAAAAAUAGAACGGAACAGAAAAAACAAAAUUUGG .....((((((...(((.((-(.....(((((((((.((((....--)))).)).)))(.....)..))))....)-))..))).-.(((((((((((..-((((((((((-(....)))))))))))..)))))))))))........))))))................. ( -55.50, z-score = -2.65, R) >consensus CCACAUGUUUCUUGGCUAAC_CCAACUGCCACCCAAGAUCGAGUC__CGAUGUUCUUGGGAUAACUUUGGCUUUCG_AUUUACCA_GUUUUUUCUGGGGGUGGGCGUGGCU_GGCCACAGUUACGCCCACCGAGAGCAAAAAAAAAAAAAAAA___CAACAAAAAAAUUGG_ ........(((((((............((((.......(((((...........)))))........))))..............................((((((((((.(....))))))))))).))))))).................................... (-21.87 = -23.59 + 1.72)
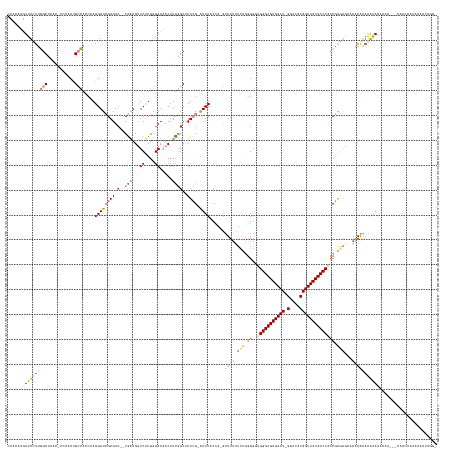
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:12:15 2011