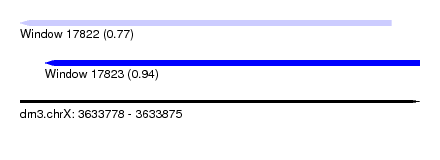

| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,633,778 – 3,633,875 |
| Length | 97 |
| Max. P | 0.940376 |

| Location | 3,633,778 – 3,633,868 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 82.83 |
| Shannon entropy | 0.27914 |
| G+C content | 0.49370 |
| Mean single sequence MFE | -32.58 |
| Consensus MFE | -21.70 |
| Energy contribution | -23.57 |
| Covariance contribution | 1.87 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.766304 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
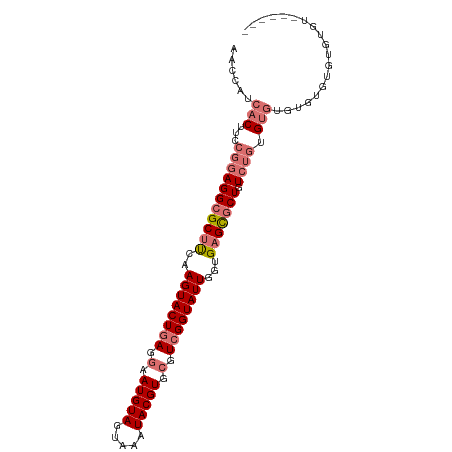
>dm3.chrX 3633778 90 - 22422827 AACCAUCACUUCCGGAGGCGCUCCAAGUACUGAGGAAUGUAGUAAAUACGUGCGUCGGUAUUGGUGAGCGCUGUCUGUGUGUGUGUGUGU---------- .((((((((...((((((((((((.((((((((.(.(((((.....))))).).)))))))).).))))))).)))).))).))).))..---------- ( -35.70, z-score = -3.51, R) >droYak2.chrX 16997370 94 + 21770863 AACCAUCACUUCCGGAGGCGCUUCAAGUACUGAGGAAUGUAGUAAAUACGUGCGUCGGUAUUGGUGAGCGCUGUCUAUGUGUGUGUGUAUGUGU------ .((((((((....((((((((((..((((((((.(.(((((.....))))).).))))))))...))))))).)))..))).))).))......------ ( -31.20, z-score = -1.97, R) >droEre2.scaffold_4690 1027284 100 - 18748788 AACCAUCACUUCCGGAGGCGCUUCAAGUACUGAGGAAUGUAGUAAAUACGUGCGUCGGUAUUGGUGAGCGCUGUCUGUGUGCGUGUAUGAGUGUAUGUGU .((((((((..((((((((((((..((((((((.(.(((((.....))))).).))))))))...))))))).)))).)...))).))).))........ ( -34.10, z-score = -1.96, R) >droAna3.scaffold_13047 83520 99 - 1816235 AACCAUCACUUCCGGAGGCGCUUCAAGUACUGAGGAAUGUAGUAAAUACGUGAGUGGGUAUUGGUGUGU-CUGUGAGUGCUACGCUGCCCUUGUGUGUGU ......((((..((((.(((((...((((((.(...(((((.....)))))...).))))))))))).)-)))..)))).(((((.......)))))... ( -29.30, z-score = -0.66, R) >consensus AACCAUCACUUCCGGAGGCGCUUCAAGUACUGAGGAAUGUAGUAAAUACGUGCGUCGGUAUUGGUGAGCGCUGUCUGUGUGUGUGUGUGUGUGU______ ......(((...(((((((((((..((((((((.(.(((((.....))))).).))))))))...))))))).)))).)))................... (-21.70 = -23.57 + 1.87)


| Location | 3,633,784 – 3,633,875 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 85.69 |
| Shannon entropy | 0.23858 |
| G+C content | 0.48702 |
| Mean single sequence MFE | -31.85 |
| Consensus MFE | -24.80 |
| Energy contribution | -26.17 |
| Covariance contribution | 1.38 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.44 |
| SVM RNA-class probability | 0.940376 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
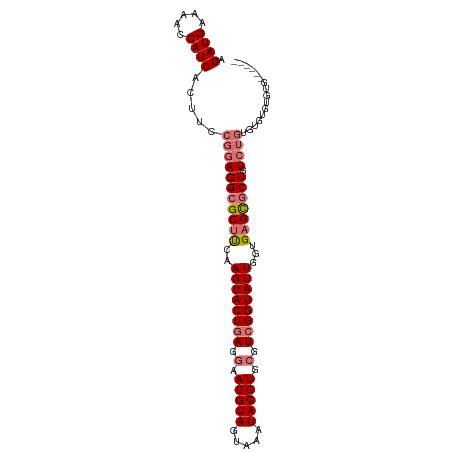
>dm3.chrX 3633784 91 - 22422827 AGAUGAAAACCAUCACUUCCGGAGGCGCUCCAAGUACUGAGGAAUGUAGUAAAUACGUGCGUCGGUAUUGGUGAGCGCUGUCUGUGUGUGU--------- .((((.....))))((...((((((((((((.((((((((.(.(((((.....))))).).)))))))).).))))))).))))...))..--------- ( -34.80, z-score = -3.63, R) >droYak2.chrX 16997377 94 + 21770863 AGAUGAAAACCAUCACUUCCGGAGGCGCUUCAAGUACUGAGGAAUGUAGUAAAUACGUGCGUCGGUAUUGGUGAGCGCUGUCUAUGUGUGUGUG------ ........((((.(((....((((((((((..((((((((.(.(((((.....))))).).))))))))...))))))).)))..))))).)).------ ( -30.50, z-score = -2.15, R) >droEre2.scaffold_4690 1027291 100 - 18748788 AGAUGAAAACCAUCACUUCCGGAGGCGCUUCAAGUACUGAGGAAUGUAGUAAAUACGUGCGUCGGUAUUGGUGAGCGCUGUCUGUGUGCGUGUAUGAGUG ........((((((((..((((((((((((..((((((((.(.(((((.....))))).).))))))))...))))))).)))).)...))).))).)). ( -34.10, z-score = -2.39, R) >droAna3.scaffold_13047 83527 99 - 1816235 AGAUGAAAACCAUCACUUCCGGAGGCGCUUCAAGUACUGAGGAAUGUAGUAAAUACGUGAGUGGGUAUUGGUGUGU-CUGUGAGUGCUACGCUGCCCUUG .((((.....))))(((..((((.(((((...((((((.(...(((((.....)))))...).))))))))))).)-)))..)))((......))..... ( -28.00, z-score = -0.79, R) >consensus AGAUGAAAACCAUCACUUCCGGAGGCGCUUCAAGUACUGAGGAAUGUAGUAAAUACGUGCGUCGGUAUUGGUGAGCGCUGUCUGUGUGUGUGUG______ .((((.....)))).....(((((((((((..((((((((.(.(((((.....))))).).))))))))...))))))).))))................ (-24.80 = -26.17 + 1.38)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:12:07 2011