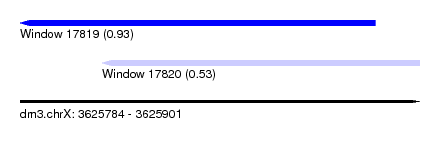

| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,625,784 – 3,625,901 |
| Length | 117 |
| Max. P | 0.927182 |

| Location | 3,625,784 – 3,625,888 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 89.42 |
| Shannon entropy | 0.15168 |
| G+C content | 0.63199 |
| Mean single sequence MFE | -35.50 |
| Consensus MFE | -34.44 |
| Energy contribution | -34.33 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.97 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.927182 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
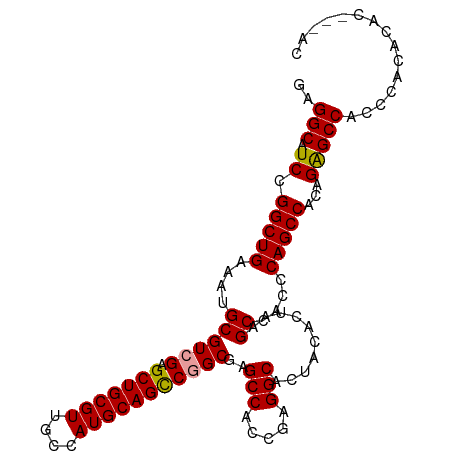
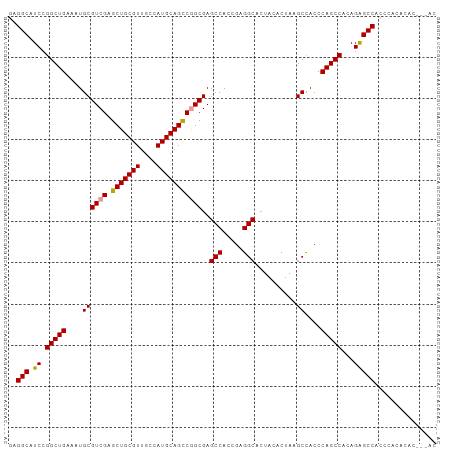
>dm3.chrX 3625784 104 - 22422827 GAGGCAUCCGGCUGAAAUGCGUCGAGCUGCGUUGCCAUGCAGUCGGCGAGCCACCGAGGCACUACACUAAGCUACCCAGCCACAGAGCCGCCCACCCACAGGAC ..(((.((.(((((....(((((((.((((((....)))))))))))..(((.....)))..........))....)))))...))))).....((....)).. ( -35.50, z-score = -0.92, R) >droEre2.scaffold_4690 1018915 104 - 18748788 GAGGCAUCCGGCUGAAAUGCGUCGAGCUGCGUUGCCAUGCAGCCGGCGAGCCACCGAGGCACUAUACUAAGCCACCCAGCCACAGAGCCACCCACACACUAAAC ..(((.((.(((((.....(((((.(((((((....)))))))))))).(((.....)))................)))))...)))))............... ( -36.70, z-score = -2.85, R) >droYak2.chrX 16989546 99 + 21770863 AAGGCAUCCGGCUGAAAUGCGUCGAGCUGCGUUGCCAUGCAGCCAGCGAGCCACGGAGGCAGUACACUAAGCCACCCAGCCACAGGGCCACCCACACAC----- ..(((....(((((....((.(((.(((((((....)))))))...)))))...((.(((((....))..))).))))))).....)))..........----- ( -34.30, z-score = -0.79, R) >consensus GAGGCAUCCGGCUGAAAUGCGUCGAGCUGCGUUGCCAUGCAGCCGGCGAGCCACCGAGGCACUACACUAAGCCACCCAGCCACAGAGCCACCCACACAC___AC ..(((.((.(((((....((((((.(((((((....)))))))))))..(((.....)))..........))....)))))...)))))............... (-34.44 = -34.33 + -0.11)

| Location | 3,625,808 – 3,625,901 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 94.98 |
| Shannon entropy | 0.06912 |
| G+C content | 0.62724 |
| Mean single sequence MFE | -35.77 |
| Consensus MFE | -35.51 |
| Energy contribution | -35.40 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -0.79 |
| Structure conservation index | 0.99 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.530288 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
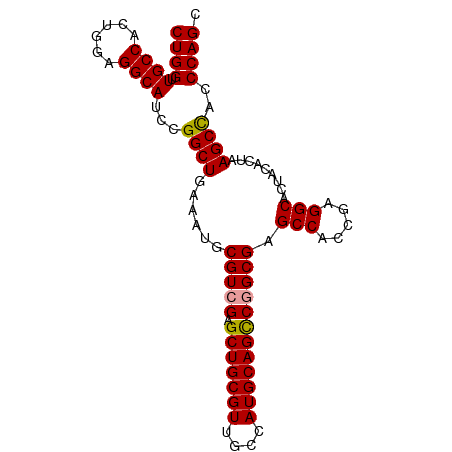
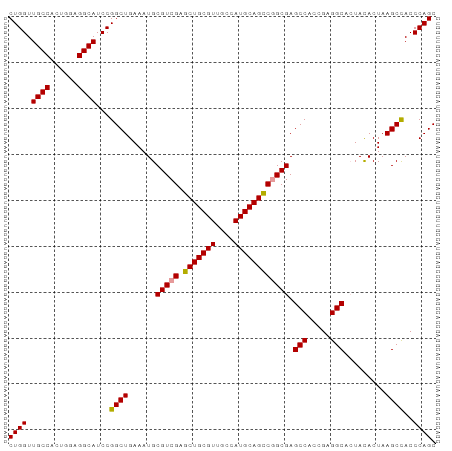
>dm3.chrX 3625808 93 - 22422827 CUGGUUGCCACUGGAGGCAUCCGGCUGAAAUGCGUCGAGCUGCGUUGCCAUGCAGUCGGCGAGCCACCGAGGCACUACACUAAGCUACCCAGC (((((((((((((.(((((.((((((((......)).))))).).)))).).)))).)))))(((.....)))...............)))). ( -34.50, z-score = -0.41, R) >droEre2.scaffold_4690 1018939 93 - 18748788 CUGGUUGCCACUGGAGGCAUCCGGCUGAAAUGCGUCGAGCUGCGUUGCCAUGCAGCCGGCGAGCCACCGAGGCACUAUACUAAGCCACCCAGC ((((.((((......))))...((((......(((((.(((((((....)))))))))))).(((.....))).........))))..)))). ( -38.50, z-score = -1.53, R) >droYak2.chrX 16989565 93 + 21770863 CUGGUUGCCACUGAAGGCAUCCGGCUGAAAUGCGUCGAGCUGCGUUGCCAUGCAGCCAGCGAGCCACGGAGGCAGUACACUAAGCCACCCAGC ((((.((((......))))...((((....(((.(((.(((((((....)))))))...)))(((.....))).))).....))))..)))). ( -34.30, z-score = -0.43, R) >consensus CUGGUUGCCACUGGAGGCAUCCGGCUGAAAUGCGUCGAGCUGCGUUGCCAUGCAGCCGGCGAGCCACCGAGGCACUACACUAAGCCACCCAGC ((((.((((......))))...((((......(((((.(((((((....)))))))))))).(((.....))).........))))..)))). (-35.51 = -35.40 + -0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:12:04 2011