
| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,567,692 – 3,567,797 |
| Length | 105 |
| Max. P | 0.850258 |

| Location | 3,567,692 – 3,567,797 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 69.80 |
| Shannon entropy | 0.54773 |
| G+C content | 0.40431 |
| Mean single sequence MFE | -20.49 |
| Consensus MFE | -10.80 |
| Energy contribution | -12.05 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.850258 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 3567692 105 - 22422827 CUGCAAG-GUUUUCUCCACUCUAUCUUCCUC-CUCUCGUUCAUC------------AUUUUGCACUUUGUUUUGCCAACGUGAGCCAAGUCGCGUUGACAAAACCCAAG-AAAAAAGUCG .((((((-(......))..............-.....(.....)------------...)))))(((.((((((.((((((((......)))))))).))))))..)))-.......... ( -20.60, z-score = -1.75, R) >droYak2.chrX 16932388 118 + 21770863 CUGCAAG-GUUUUCCCCACUUUAUCUUCCUC-UUCUCAUUCAUCCGCCUCUUCUUUGUUUUGCACCUUGUUUUGCCAAUGUGAGCCAAGUCGCGUUGACAAAAGCAAAGAAAAAAAGUUG ......(-(......))..............-..................((((((((((((((........)))((((((((......))))))))...)))))))))))......... ( -20.50, z-score = -0.52, R) >droSec1.super_4 3050051 105 + 6179234 CUGCAAG-GUUUUCUCCACUGUAUCUUCCUC-CUCUCGUUCAUC------------AUUUUGCACUUUGUUUUGUCAACGAGAGCCAAGUCGCGUUGACAAAAGCCAAG-AAAAAAGUCG ......(-(((((...((.(((..(((....-((((((((...(------------(....((.....))..))..))))))))..)))..))).))...))))))...-.......... ( -19.90, z-score = -0.49, R) >droSim1.chrX 2608275 105 - 17042790 CUGCAAG-GUUUUCUCCACUCUAUCUCCCUC-CUCUCGUUCAGC------------AUUUUGCACUUUGUUUUGUCAACGUGAGCCAAGUCGCGUUGACAAAAGCCAAG-AAAAAAGUCG .((((((-(......))..............-.....(.....)------------...)))))(((.(((((((((((((((......)))))))))).))))).)))-.......... ( -24.40, z-score = -2.17, R) >droEre2.scaffold_4690 963608 119 - 18748788 CUGCAAGCUUUUUUUCCACUUUAUCUUCCUC-UUCUCGUUCAUCAUCCUCUUCUUUGUUUUGCACUUUGUUUUGUCAACGUGAGCCAAGUCGCGUUCACAAAAUCAAAGAAAAAGAGUUG .....(((((((((((...............-.....((....((..........))....))..((((((((((.(((((((......))))))).)))))).))))))))))))))). ( -24.20, z-score = -2.83, R) >droVir3.scaffold_12728 19383 88 - 808307 AAGCUCA-GUUUGCUCGAAUUAAUUUUUUUCAAUCUCGUUU---------------AAUCGAAAUAUUUUUAAGUCGA--UAAAUUGAAUUCCAAUGGAAAUGGUA-------------- ....(((-(((((.((((.((((....((((..........---------------....)))).....)))).))))--)))))))).((((...))))......-------------- ( -13.34, z-score = -0.29, R) >consensus CUGCAAG_GUUUUCUCCACUUUAUCUUCCUC_CUCUCGUUCAUC____________AUUUUGCACUUUGUUUUGUCAACGUGAGCCAAGUCGCGUUGACAAAAGCAAAG_AAAAAAGUCG .....................................................................((((((((((((((......))))))))))))))................. (-10.80 = -12.05 + 1.25)

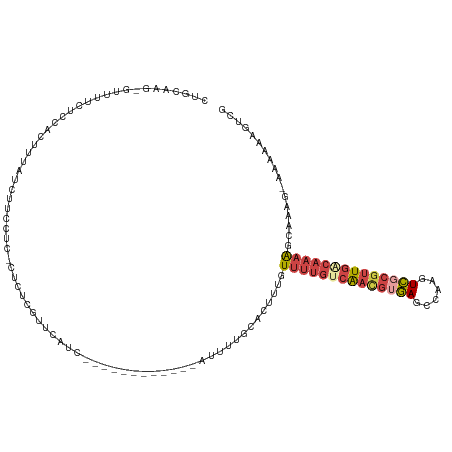
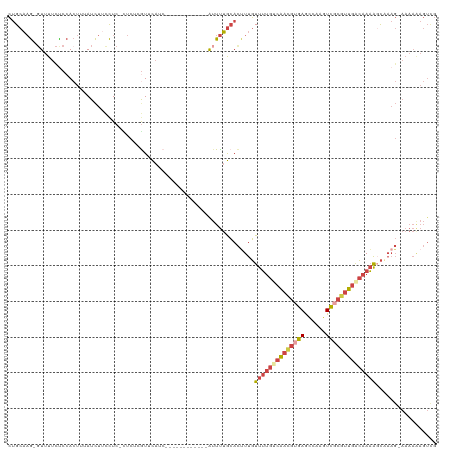
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:11:54 2011