
| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,527,146 – 3,527,207 |
| Length | 61 |
| Max. P | 0.537965 |

| Location | 3,527,146 – 3,527,207 |
|---|---|
| Length | 61 |
| Sequences | 5 |
| Columns | 62 |
| Reading direction | forward |
| Mean pairwise identity | 62.58 |
| Shannon entropy | 0.67102 |
| G+C content | 0.43915 |
| Mean single sequence MFE | -13.04 |
| Consensus MFE | -4.88 |
| Energy contribution | -5.36 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.537965 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
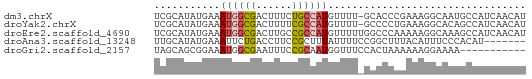
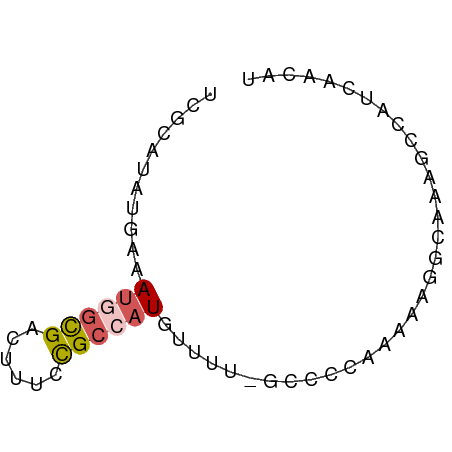
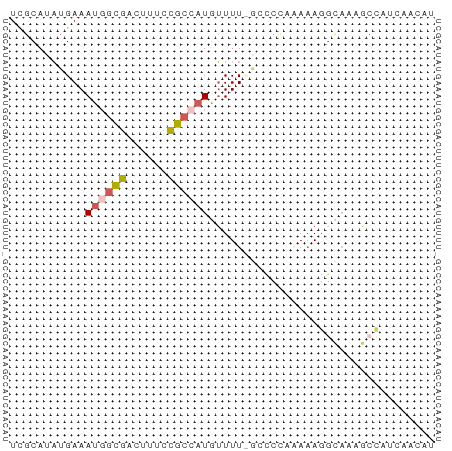
>dm3.chrX 3527146 61 + 22422827 UCGCAUAUGAAAUGGCGACUUUCUGCCAUGUUUU-GCACCCGAAAGGCAAUGCCAUCAACAU ..(((...((.((((((......)))))).)).)-))..((....))............... ( -14.40, z-score = -0.95, R) >droYak2.chrX 16889738 61 - 21770863 UCGCAUAUGAAAUGGCGACUUUUCGCCAUGUUUU-GCCCCUGAAAGGCACAGCCAUCAACAU .......(((.(((((((....)))))))(((.(-(((.......)))).)))..))).... ( -17.00, z-score = -2.15, R) >droEre2.scaffold_4690 920477 62 + 18748788 UCGCAUAUGAAAUGGCGACUUGCCGCCAUGUUUUUGGCCCAAAAAGGCAAAGCCAUCAACAU .......(((..((((...(((((((((......)))).......))))).))))))).... ( -17.41, z-score = -1.43, R) >droAna3.scaffold_13248 3869981 55 + 4840945 UUGCAUAUGAAAUUCUGACCUUCCGCUUUAUUUUCCGGCUUUACAUUUCCCACAU------- ........(((((..(((....(((..........)))..))).)))))......------- ( -1.20, z-score = 1.37, R) >droGri2.scaffold_2157 489 51 - 1193 UAGCAGCGGAAAUGGCGAAUUUCCGCAAUGGUUUCCACUAAAAAAGGAAAA----------- .....((((((((.....)))))))).....(((((.........))))).----------- ( -15.20, z-score = -2.77, R) >consensus UCGCAUAUGAAAUGGCGACUUUCCGCCAUGUUUU_GCCCCAAAAAGGCAAAGCCAUCAACAU ...........((((((......))))))................................. ( -4.88 = -5.36 + 0.48)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:11:48 2011