
| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,484,418 – 3,484,522 |
| Length | 104 |
| Max. P | 0.989888 |

| Location | 3,484,418 – 3,484,522 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.06 |
| Shannon entropy | 0.39795 |
| G+C content | 0.38299 |
| Mean single sequence MFE | -24.34 |
| Consensus MFE | -11.24 |
| Energy contribution | -13.00 |
| Covariance contribution | 1.76 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.627754 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
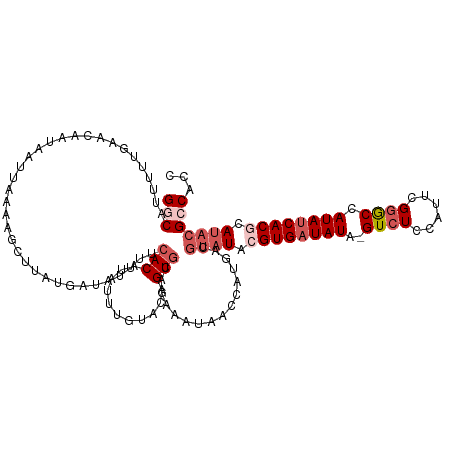
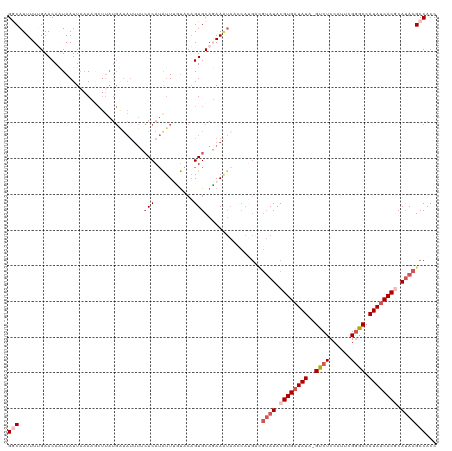
>dm3.chrX 3484418 104 + 22422827 GGCAUUUUUGAACAAUAAUUACAC---------------ACAGAUUUUUCCAGUGCAGAAAUAAGCAUGACGUAUACGUGAUAUA-GUCUCCAUUUGGACCAUAUCACACAUACGCCACC (((.....................---------------.............((((........))))...((((..(((((((.-((((......)))).)))))))..)))))))... ( -23.70, z-score = -3.49, R) >droEre2.scaffold_4690 899016 107 + 18748788 GACAUUUUGGAACACUAAUUAAAUGCUUAUGGUAUUUUCACAUACUUCUCCUGUGCAGAA-------------AUACGUGUUAUACGUCUCCAUUUGAGCCAUAUCACGCAUACGCCACC .......(((.....................(((((((..((((.......))))..)))-------------))))((((.(((.(.(((.....))).).))).)))).....))).. ( -16.10, z-score = 0.05, R) >droYak2.chrX 16868190 119 - 21770863 GACAUUUUUGAACACUAAUUUAACUCUUAUGAUAUUUUCACAUAUUUUUGCAGUGCAGAAAUAACCAUGAUGUAUACGUGAUAUA-GUCUCCAUUCGGACCAUAUCACACAUACGCCACC .(((((..((...................(((.....)))..(((((((((...)))))))))..)).)))))....(((((((.-((((......)))).)))))))............ ( -22.80, z-score = -2.68, R) >droSec1.super_4 2989645 118 - 6179234 GGCAUUUU-GAACAAUAAUUAAAAGCUUAUGAUAUUUGCACAUAUUGGUACAGUGCAUAAGUAACCAAGUCGUAUACGUGAUAUA-GUCUCCAAUCGGGCCAUAUCACGCACACGCCACC (((.....-...............(((((((.(((((((........))).))))))))))).........((...((((((((.-((((......)))).))))))))...)))))... ( -28.20, z-score = -1.81, R) >droSim1.chrX 2553095 119 + 17042790 GGCAUUUUUGAACAAUAAUUAAAAGCUUAUGAUAUUUACACAUGUUGGCACAGUGCAUAAGUAACCAUGACGUAUACGUGAUAUA-GUCUCCAAUCGGGCCAUAUCACGCAUACGCCACC (((.....................(((((((.......(((.(((....))))))))))))).........((((.((((((((.-((((......)))).)))))))).)))))))... ( -30.91, z-score = -2.51, R) >consensus GGCAUUUUUGAACAAUAAUUAAAAGCUUAUGAUAUUUUCACAUAUUUGUACAGUGCAGAAAUAACCAUGACGUAUACGUGAUAUA_GUCUCCAUUCGGGCCAUAUCACGCAUACGCCACC (((...................................(((...........)))................((((.(((((((...((((......))))..))))))).)))))))... (-11.24 = -13.00 + 1.76)

| Location | 3,484,418 – 3,484,522 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.06 |
| Shannon entropy | 0.39795 |
| G+C content | 0.38299 |
| Mean single sequence MFE | -33.74 |
| Consensus MFE | -18.04 |
| Energy contribution | -19.44 |
| Covariance contribution | 1.40 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.93 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.39 |
| SVM RNA-class probability | 0.989888 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
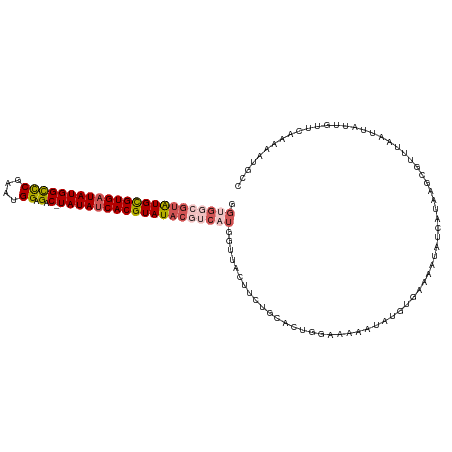
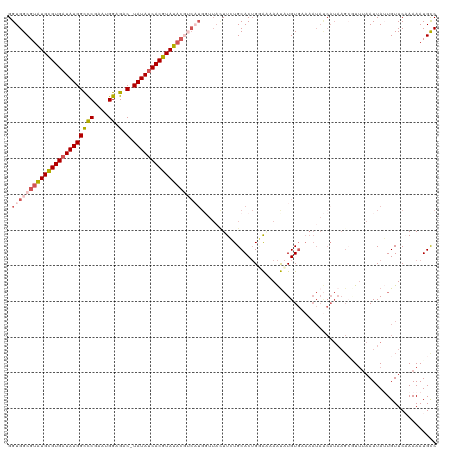
>dm3.chrX 3484418 104 - 22422827 GGUGGCGUAUGUGUGAUAUGGUCCAAAUGGAGAC-UAUAUCACGUAUACGUCAUGCUUAUUUCUGCACUGGAAAAAUCUGU---------------GUGUAAUUAUUGUUCAAAAAUGCC .((((((((((((((((((((((........)))-))))))))))))))))))).........(((((((((....)))).---------------)))))................... ( -34.90, z-score = -4.51, R) >droEre2.scaffold_4690 899016 107 - 18748788 GGUGGCGUAUGCGUGAUAUGGCUCAAAUGGAGACGUAUAACACGUAU-------------UUCUGCACAGGAGAAGUAUGUGAAAAUACCAUAAGCAUUUAAUUAGUGUUCCAAAAUGUC (((((((....))).(((((.(((.....))).)))))..(((((((-------------((((.(...).)))))))))))....))))...((((((.....)))))).......... ( -26.60, z-score = -1.34, R) >droYak2.chrX 16868190 119 + 21770863 GGUGGCGUAUGUGUGAUAUGGUCCGAAUGGAGAC-UAUAUCACGUAUACAUCAUGGUUAUUUCUGCACUGCAAAAAUAUGUGAAAAUAUCAUAAGAGUUAAAUUAGUGUUCAAAAAUGUC .(((..((((((((((((((((((.....).)))-))))))))))))))..))).....(((..((((((....(((.(((((.....)))))...)))....))))))..)))...... ( -30.30, z-score = -2.32, R) >droSec1.super_4 2989645 118 + 6179234 GGUGGCGUGUGCGUGAUAUGGCCCGAUUGGAGAC-UAUAUCACGUAUACGACUUGGUUACUUAUGCACUGUACCAAUAUGUGCAAAUAUCAUAAGCUUUUAAUUAUUGUUC-AAAAUGCC ...(((((((((((((((((((((....)).).)-))))))))))))))(((.((((((((((((...(((((......))))).....))))))....))))))..))).-.....))) ( -36.60, z-score = -2.79, R) >droSim1.chrX 2553095 119 - 17042790 GGUGGCGUAUGCGUGAUAUGGCCCGAUUGGAGAC-UAUAUCACGUAUACGUCAUGGUUACUUAUGCACUGUGCCAACAUGUGUAAAUAUCAUAAGCUUUUAAUUAUUGUUCAAAAAUGCC ((((((((((((((((((((((((....)).).)-)))))))))))))))))(((((...(((((((.(((....))))))))))..))))).........................))) ( -40.30, z-score = -3.66, R) >consensus GGUGGCGUAUGCGUGAUAUGGCCCGAAUGGAGAC_UAUAUCACGUAUACGUCAUGGUUACUUCUGCACUGGAAAAAUAUGUGAAAAUAUCAUAAGCGUUUAAUUAUUGUUCAAAAAUGCC .(((((((((((((((((((.(((....)).)...))))))))))))))))))).................................................................. (-18.04 = -19.44 + 1.40)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:11:46 2011