
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 9,877,259 – 9,877,358 |
| Length | 99 |
| Max. P | 0.987378 |

| Location | 9,877,259 – 9,877,358 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 86.97 |
| Shannon entropy | 0.24526 |
| G+C content | 0.32955 |
| Mean single sequence MFE | -23.90 |
| Consensus MFE | -17.56 |
| Energy contribution | -17.70 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.94 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.28 |
| SVM RNA-class probability | 0.987378 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
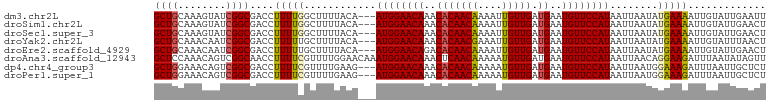
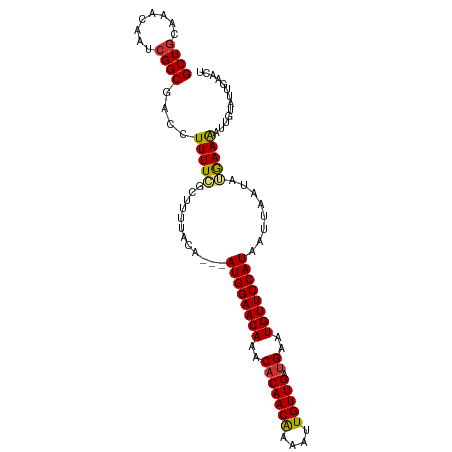
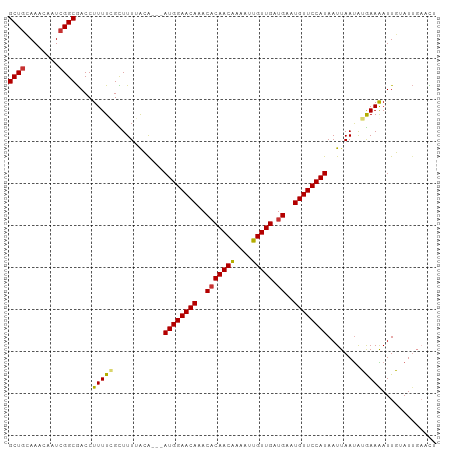
>dm3.chr2L 9877259 99 + 23011544 GCUGCAAAGUAUCGGCGACCUUUUGGCUUUUACA---AUGGAACAAACACAACAAAAUUGUUGAUGAAUGUUCCAUAAUUAAUAUGAAAAUUGUAUUGAAUU ..(((((.(((..(((.(.....).)))..))).---((((((((..(((((((....))))).))..))))))))..............)))))....... ( -22.50, z-score = -2.41, R) >droSim1.chr2L 9661806 99 + 22036055 GCUGCAAAGUAUCGGCGACCUUUUGGCUUUUACA---AUGGAACAAACACAACAAAAUUGUUGAUGAAUGUUCCAUAAUUAAUAUGAAAAUUGUAUUGAACU ..(((((.(((..(((.(.....).)))..))).---((((((((..(((((((....))))).))..))))))))..............)))))....... ( -22.50, z-score = -2.59, R) >droSec1.super_3 5323315 99 + 7220098 GCUGCAAAGUAUCGGCGACCUUUUGGCUUUUACA---AUGGAACAAACACAACAAAAUUGUUGAUGAAUGUUCCAUAAUUAAUAUGAAAAUUGUAUUGAACU ..(((((.(((..(((.(.....).)))..))).---((((((((..(((((((....))))).))..))))))))..............)))))....... ( -22.50, z-score = -2.59, R) >droYak2.chr2L 12566955 99 + 22324452 GCUGCAAACAAUCGGCGACCUUUUUGCUUUUACA---AUGGAACAAACACAACGAAAUUGUUGAUGAAUGUUCCAUAAUUAAUAUGAAAAUUGUAUUUAACU ..(((((......(((((.....)))))......---((((((((..(((((((....))))).))..))))))))..............)))))....... ( -19.70, z-score = -2.37, R) >droEre2.scaffold_4929 10491026 99 + 26641161 GCUGCAAACAAUCGGCGACCUUUUUGCUUUUACA---AUGGAACAGACACAACAAAAUUGUUGAUGAAUGUUCCAUAAUUAAUAUGAAAAUUGUAUUGAACU ((((........))))..................---((((((((..(((((((....))))).))..))))))))..((((((..(...)..))))))... ( -22.20, z-score = -2.92, R) >droAna3.scaffold_12943 1322751 102 + 5039921 GCUCCAAACAGUCGGCAACCUUUUCGUUUUGGAACAAAUGGAACAAACUCAACAAAAAUGUUGAUGAAUGUUCCAUAAUUAACAGGAAGAUUUAAUAUAGUU (.((((((....(((........))).)))))).)..((((((((..(((((((....)))))).)..)))))))).......................... ( -20.60, z-score = -1.51, R) >dp4.chr4_group3 11400073 99 - 11692001 GCUGGAAACAGUCGGCGACCUUUUCGUUUUGAAG---AUGGAACAAACACAACAAAAAUGUUGAUGAAUGUUCCAUAAUUAAUGGAAAGAUUUAAUUGCUCU ((((....)))).(((((.(((((((((......---((((((((..(((((((....))))).))..))))))))....)))))))))......))))).. ( -30.60, z-score = -4.58, R) >droPer1.super_1 8516588 99 - 10282868 GCUGGAAACAGUCGGCGACCUUUUCGUUUUGAAG---AUGGAACAAACACAACAAAAAUGUUGAUGAAUGUUCCAUAAUUAAUGGAAAGAUUUAAUUGCUCU ((((....)))).(((((.(((((((((......---((((((((..(((((((....))))).))..))))))))....)))))))))......))))).. ( -30.60, z-score = -4.58, R) >consensus GCUGCAAACAAUCGGCGACCUUUUCGCUUUUACA___AUGGAACAAACACAACAAAAUUGUUGAUGAAUGUUCCAUAAUUAAUAUGAAAAUUGUAUUGAACU ((((........)))).....................((((((((..(((((((....))))).))..)))))))).......................... (-17.56 = -17.70 + 0.14)
| Location | 9,877,259 – 9,877,358 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 86.97 |
| Shannon entropy | 0.24526 |
| G+C content | 0.32955 |
| Mean single sequence MFE | -21.76 |
| Consensus MFE | -16.63 |
| Energy contribution | -16.73 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.948391 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 9877259 99 - 23011544 AAUUCAAUACAAUUUUCAUAUUAAUUAUGGAACAUUCAUCAACAAUUUUGUUGUGUUUGUUCCAU---UGUAAAAGCCAAAAGGUCGCCGAUACUUUGCAGC .........(((...((...(((...((((((((..((.(((((....)))))))..))))))))---..)))..(((....)))....))....))).... ( -20.10, z-score = -2.04, R) >droSim1.chr2L 9661806 99 - 22036055 AGUUCAAUACAAUUUUCAUAUUAAUUAUGGAACAUUCAUCAACAAUUUUGUUGUGUUUGUUCCAU---UGUAAAAGCCAAAAGGUCGCCGAUACUUUGCAGC .........(((...((...(((...((((((((..((.(((((....)))))))..))))))))---..)))..(((....)))....))....))).... ( -20.10, z-score = -1.91, R) >droSec1.super_3 5323315 99 - 7220098 AGUUCAAUACAAUUUUCAUAUUAAUUAUGGAACAUUCAUCAACAAUUUUGUUGUGUUUGUUCCAU---UGUAAAAGCCAAAAGGUCGCCGAUACUUUGCAGC .........(((...((...(((...((((((((..((.(((((....)))))))..))))))))---..)))..(((....)))....))....))).... ( -20.10, z-score = -1.91, R) >droYak2.chr2L 12566955 99 - 22324452 AGUUAAAUACAAUUUUCAUAUUAAUUAUGGAACAUUCAUCAACAAUUUCGUUGUGUUUGUUCCAU---UGUAAAAGCAAAAAGGUCGCCGAUUGUUUGCAGC ....................(((...((((((((..((.((((......))))))..))))))))---..)))..(((((...(((...)))..)))))... ( -21.50, z-score = -2.39, R) >droEre2.scaffold_4929 10491026 99 - 26641161 AGUUCAAUACAAUUUUCAUAUUAAUUAUGGAACAUUCAUCAACAAUUUUGUUGUGUCUGUUCCAU---UGUAAAAGCAAAAAGGUCGCCGAUUGUUUGCAGC ....................(((...((((((((..((.(((((....)))))))..))))))))---..)))..(((((...(((...)))..)))))... ( -21.40, z-score = -2.11, R) >droAna3.scaffold_12943 1322751 102 - 5039921 AACUAUAUUAAAUCUUCCUGUUAAUUAUGGAACAUUCAUCAACAUUUUUGUUGAGUUUGUUCCAUUUGUUCCAAAACGAAAAGGUUGCCGACUGUUUGGAGC ..........................((((((((..(.((((((....)))))))..))))))))..((((((((.......((...)).....)))))))) ( -25.50, z-score = -2.83, R) >dp4.chr4_group3 11400073 99 + 11692001 AGAGCAAUUAAAUCUUUCCAUUAAUUAUGGAACAUUCAUCAACAUUUUUGUUGUGUUUGUUCCAU---CUUCAAAACGAAAAGGUCGCCGACUGUUUCCAGC ...((.....................((((((((..((.(((((....)))))))..))))))))---.........((((.((((...)))).))))..)) ( -22.70, z-score = -2.79, R) >droPer1.super_1 8516588 99 + 10282868 AGAGCAAUUAAAUCUUUCCAUUAAUUAUGGAACAUUCAUCAACAUUUUUGUUGUGUUUGUUCCAU---CUUCAAAACGAAAAGGUCGCCGACUGUUUCCAGC ...((.....................((((((((..((.(((((....)))))))..))))))))---.........((((.((((...)))).))))..)) ( -22.70, z-score = -2.79, R) >consensus AGUUCAAUACAAUUUUCAUAUUAAUUAUGGAACAUUCAUCAACAAUUUUGUUGUGUUUGUUCCAU___UGUAAAAGCCAAAAGGUCGCCGAUUGUUUGCAGC ..........................((((((((..((.(((((....)))))))..))))))))............((((..(((...)))..)))).... (-16.63 = -16.73 + 0.10)

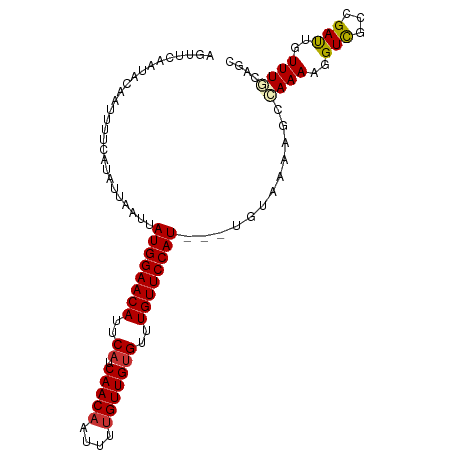
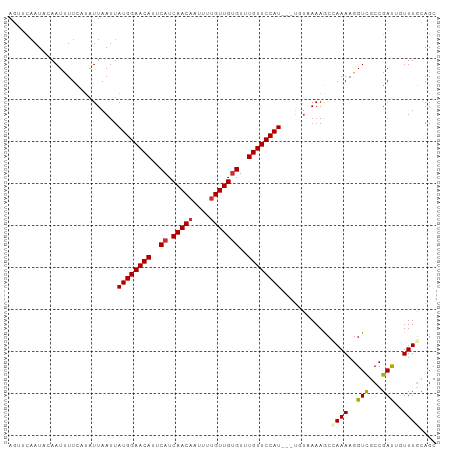
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:29:09 2011