
| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,276,058 – 3,276,118 |
| Length | 60 |
| Max. P | 0.699163 |

| Location | 3,276,058 – 3,276,118 |
|---|---|
| Length | 60 |
| Sequences | 9 |
| Columns | 70 |
| Reading direction | reverse |
| Mean pairwise identity | 74.65 |
| Shannon entropy | 0.47818 |
| G+C content | 0.42867 |
| Mean single sequence MFE | -13.14 |
| Consensus MFE | -7.08 |
| Energy contribution | -7.36 |
| Covariance contribution | 0.27 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.699163 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 3276058 60 - 22422827 -------GACGACAGC--AGCAACUUUUGUUGCUAAGCGAACGAAGCCAAGAUCAAAUCGUCACAAUAA- -------(((((..((--((((((....))))))..))((.(........).))...))))).......- ( -15.80, z-score = -2.84, R) >droAna3.scaffold_12613 430824 56 + 519072 ----------GACGG---AACAACUUUUGUUGCUAAGCGAACGAAGCCAAGAUCAAAUCGUAAUAAUAA- ----------.((((---..((((....))))((..((.......))..))......))))........- ( -6.40, z-score = 0.13, R) >droEre2.scaffold_4690 688450 60 - 18748788 -------GACGGCAGC--AGCAACUUUUGUUGCUAAGCCAACGAAGCCAAGAUCAAAUCGUAAUAAUAA- -------...((((((--((((.....)))))))..))).((((.............))))........- ( -13.92, z-score = -2.02, R) >droYak2.chrX 16669527 60 + 21770863 -------GACGGCAGC--AGCAACUUUUGUUGCUAAGCGAACGAAGCCAAGAUCAAAUCGUAAUAAUAA- -------...(((.((--((((((....))))))..)).......))).....................- ( -11.70, z-score = -0.70, R) >droSec1.super_10 2981071 60 - 3036183 -------GACGGCAGC--AGCAACUUUUGUUGCUAAGCGAACGAAGCCAAGAUCAAAUCGUAAUAAUAA- -------...(((.((--((((((....))))))..)).......))).....................- ( -11.70, z-score = -0.70, R) >droSim1.chrX 2362086 60 - 17042790 -------GACGGCAGC--AGCAACUUUUGUUGCUAAGCGAACGAAGCCAAGAUCAAAUCGUAAUAAUAA- -------...(((.((--((((((....))))))..)).......))).....................- ( -11.70, z-score = -0.70, R) >droWil1.scaffold_181096 1889789 54 + 12416693 -------GAUAAAAGC--AAAAACUUUUGUUGCUAAGCA---GCAGCAGCAGUGAGACAACGACGA---- -------.........--....(((.((((((((....)---))))))).))).............---- ( -11.20, z-score = -0.63, R) >droVir3.scaffold_12970 3718588 55 + 11907090 ---------CGACAGCUUGGCAACUUUUGUUGCUAAGCG-----CGCCAAGAUCAAAUCAAGACAACAG- ---------((...((((((((((....)))))))))).-----)).......................- ( -16.80, z-score = -2.83, R) >droMoj3.scaffold_6482 1236959 64 - 2735782 AGCGAUAACCGACAGCUUGGCAACUUUUGUUGCUAAGCG-----CGCCAAGAUCAAAUCAAGAC-ACAGC ...(((...((...((((((((((....)))))))))).-----)).....)))..........-..... ( -19.00, z-score = -2.50, R) >consensus _______GACGACAGC__AGCAACUUUUGUUGCUAAGCGAACGAAGCCAAGAUCAAAUCGUAAUAAUAA_ ..............((..((((((....))))))..))................................ ( -7.08 = -7.36 + 0.27)

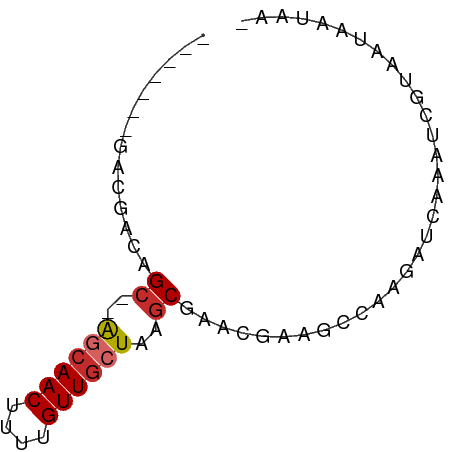
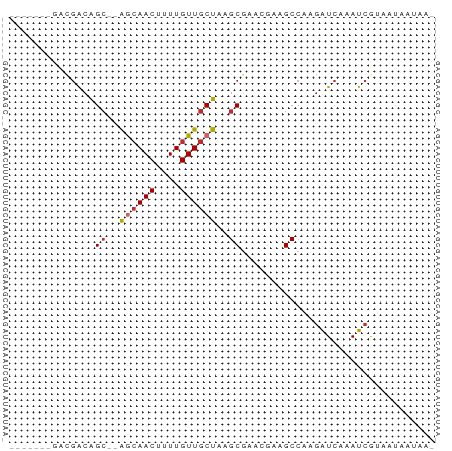
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:11:21 2011