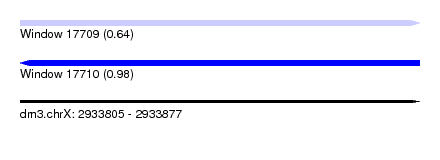

| Sequence ID | dm3.chrX |
|---|---|
| Location | 2,933,805 – 2,933,877 |
| Length | 72 |
| Max. P | 0.976742 |

| Location | 2,933,805 – 2,933,877 |
|---|---|
| Length | 72 |
| Sequences | 11 |
| Columns | 72 |
| Reading direction | forward |
| Mean pairwise identity | 94.49 |
| Shannon entropy | 0.12835 |
| G+C content | 0.50273 |
| Mean single sequence MFE | -19.90 |
| Consensus MFE | -18.84 |
| Energy contribution | -19.22 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.637344 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
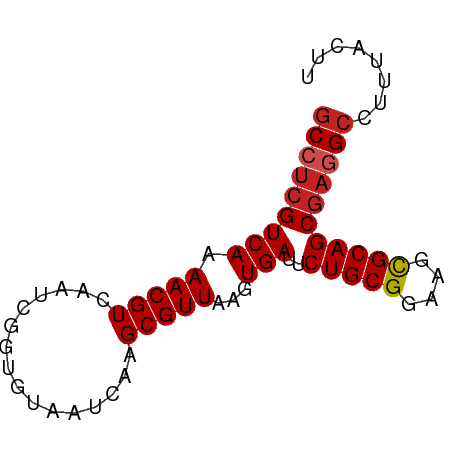
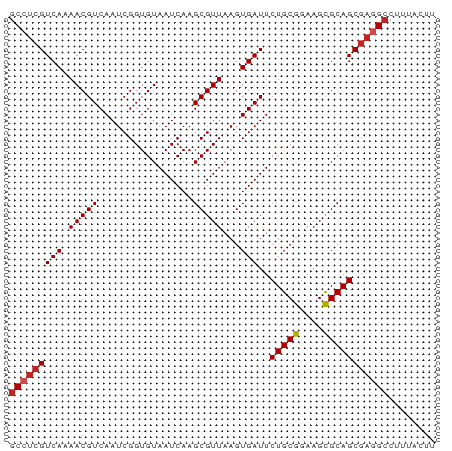
>dm3.chrX 2933805 72 + 22422827 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUUACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ ( -21.39, z-score = -1.40, R) >droSim1.chrX 2072570 72 + 17042790 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUUACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ ( -21.39, z-score = -1.40, R) >droSec1.super_10 2673686 72 + 3036183 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUUACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ ( -21.39, z-score = -1.40, R) >droYak2.chrX 16341049 72 - 21770863 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUUACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ ( -21.39, z-score = -1.40, R) >droEre2.scaffold_4690 361531 72 + 18748788 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUUACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ ( -21.39, z-score = -1.40, R) >droAna3.scaffold_12613 2246 72 - 519072 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUAACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ ( -21.39, z-score = -1.49, R) >dp4.chrXL_group1e 2412553 72 + 12523060 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUUACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ ( -21.39, z-score = -1.40, R) >droPer1.super_18 1282218 72 + 1952607 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUUACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ ( -21.39, z-score = -1.40, R) >droWil1.scaffold_181096 10072705 72 - 12416693 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAAGGCCUUAAAC (((((((((.(((((................)))))...)))..(((((....))))))).))))....... ( -17.59, z-score = -0.40, R) >droVir3.scaffold_12726 274775 65 + 2840439 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCAUAGCC------- ((...((((.(((((................)))))...)))).(((((....)))))....)).------- ( -12.39, z-score = 0.63, R) >droGri2.scaffold_15203 2488335 67 + 11997470 GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGUGCAGCGAGGCCUG----- (((((((((.(((((................)))))...)))..(((((....)))))))))))...----- ( -17.79, z-score = -0.71, R) >consensus GCCUCGUCAAAACGUCAAUCGGUGUAAUCAAGCGUUAAGUGAUUCUGCGGAAGCGCAGCGAGGCCUUUACUU (((((((((.(((((................)))))...)))..(((((....)))))))))))........ (-18.84 = -19.22 + 0.37)

| Location | 2,933,805 – 2,933,877 |
|---|---|
| Length | 72 |
| Sequences | 11 |
| Columns | 72 |
| Reading direction | reverse |
| Mean pairwise identity | 94.49 |
| Shannon entropy | 0.12835 |
| G+C content | 0.50273 |
| Mean single sequence MFE | -20.36 |
| Consensus MFE | -20.07 |
| Energy contribution | -20.11 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.99 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.976742 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
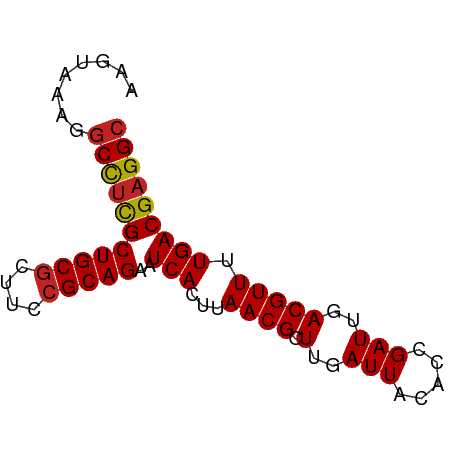
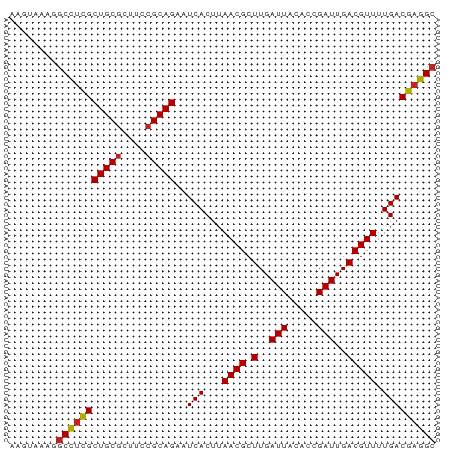
>dm3.chrX 2933805 72 - 22422827 AAGUAAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -21.60, z-score = -2.08, R) >droSim1.chrX 2072570 72 - 17042790 AAGUAAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -21.60, z-score = -2.08, R) >droSec1.super_10 2673686 72 - 3036183 AAGUAAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -21.60, z-score = -2.08, R) >droYak2.chrX 16341049 72 + 21770863 AAGUAAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -21.60, z-score = -2.08, R) >droEre2.scaffold_4690 361531 72 - 18748788 AAGUAAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -21.60, z-score = -2.08, R) >droAna3.scaffold_12613 2246 72 + 519072 AAGUUAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -21.60, z-score = -2.11, R) >dp4.chrXL_group1e 2412553 72 - 12523060 AAGUAAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -21.60, z-score = -2.08, R) >droPer1.super_18 1282218 72 - 1952607 AAGUAAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -21.60, z-score = -2.08, R) >droWil1.scaffold_181096 10072705 72 + 12416693 GUUUAAGGCCUUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC .......((((.(((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) ( -17.80, z-score = -0.97, R) >droVir3.scaffold_12726 274775 65 - 2840439 -------GGCUAUGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC -------.(((.(((((((....)))))..(((...((((.(..(((.....)))..))))).))))).))) ( -13.20, z-score = -0.16, R) >droGri2.scaffold_15203 2488335 67 - 11997470 -----CAGGCCUCGCUGCACUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC -----...((((((((((......))))..(((...((((.(..(((.....)))..))))).))))))))) ( -20.20, z-score = -2.99, R) >consensus AAGUAAAGGCCUCGCUGCGCUUCCGCAGAAUCACUUAACGCUUGAUUACACCGAUUGACGUUUUGACGAGGC ........(((((((((((....)))))..(((...((((.(..(((.....)))..))))).))))))))) (-20.07 = -20.11 + 0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:10:34 2011