
| Sequence ID | dm3.chrX |
|---|---|
| Location | 2,774,339 – 2,774,435 |
| Length | 96 |
| Max. P | 0.627542 |

| Location | 2,774,339 – 2,774,435 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 63.71 |
| Shannon entropy | 0.62857 |
| G+C content | 0.54949 |
| Mean single sequence MFE | -36.27 |
| Consensus MFE | -11.83 |
| Energy contribution | -12.35 |
| Covariance contribution | 0.51 |
| Combinations/Pair | 1.47 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.627542 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 2774339 96 - 22422827 GGAUGCCUCUUGCGGUGGCC--GAGGACAAUGGCCAAGUCAUAAACGGCGACUGCAGGAGGCAUUUAAGCC-----CCACAUUGUCGCGGUCAACGGCAAAAA---------------- ((((((((((((((((.(((--(..(((.........))).....)))).)))))))))))))))).....-----.....((((((.......))))))...---------------- ( -42.70, z-score = -3.79, R) >droSec1.super_10 2515676 96 - 3036183 GGAUGGCUCUUGCGGUGGCC--GAGGACAAUGGCCAAGUCAUAAACGGCGACUGCAGGAGGCAUUUAAGCC-----CCACAUCGUCGCGGUCAACGGCAAAAA---------------- (((((.((((((((((.(((--(..(((.........))).....)))).)))))))))).)))))..(((-----....((((...))))....))).....---------------- ( -38.10, z-score = -2.40, R) >droYak2.chrX 16184184 96 + 21770863 GGAAGGCUUUUGCGGUGACC--GAGGACAAUGGCCAAGUCAUAAACGGUGACUGCAGCAGGCAUUUAAGCC-----CCACAACGUCGCGGUCAACGGCAAAAA---------------- .......((((((.((((((--(..(((....((..((((((.....))))))...)).(((......)))-----.......))).))))).)).)))))).---------------- ( -34.20, z-score = -1.98, R) >droEre2.scaffold_4690 195127 96 - 18748788 UGCAGGCUUUUGCGGUUGCC--GAGGACAAUGGCCAAGUCAUAAGCGGCGAAUGCAGCAGGCAUUUAAGCC-----CCACAACGUCGCGGUCAAUGGCAAAAA---------------- .......((((((.((((((--(..(((.(((((...)))))..(.(((((((((.....))))))..)))-----.).....))).))).)))).)))))).---------------- ( -31.90, z-score = -0.45, R) >droAna3.scaffold_13248 4613900 98 + 4840945 ----GGACUUUGCGGUGGCCGAGAGGACAAUGGCCAAGUCAUAAACUGCCACUGCAGGAGGCAUUUAGCCCAGAGUUGAUGUCCUCGCGGCGGACGGCAAAA----------------- ----....(((((.((.((((.(((((((.....(((.((.....((((....))))(.(((.....)))).)).))).))))))).)))).).).))))).----------------- ( -39.30, z-score = -2.31, R) >dp4.chrXL_group1e 2214150 116 - 12523060 -GGAAAUGCUUGCGGUGGCC--CAGGACAAUGGCCAACUCAUAAACUGUUCCCGCUGAAUCAACUGGAACGAGAGCUGCGGGAGCAGACUGUACUAGUGGAACCUGGCUAACAGCGAAA -........((((..(((((--.(((.......(((....(((..((((((((((.(..((.........))...).))))))))))..))).....)))..))))))))...)))).. ( -36.40, z-score = -1.01, R) >droPer1.super_18 1085088 115 - 1952607 -GGAAAUGCUUGCGGUGGCC--CAGGACAAUGGCCAACUCAUAAACUGUUCCCGCUGAAUCAACUGGAACGAGAGCUGC-GGAGCAGACUGUACUAGUGGAACCUGGUUAACAGCGAAA -.....((((((((((((((--.........))))..........(((((((.(.((...)).).))))).)).)))))-.)))))(.(((((((((......)))))..))))).... ( -31.30, z-score = 0.14, R) >consensus _GAAGGCUCUUGCGGUGGCC__GAGGACAAUGGCCAAGUCAUAAACGGCGACUGCAGGAGGCAUUUAAGCC_____CCACAUCGUCGCGGUCAACGGCAAAAA________________ .........((((((((((......(((.........))).......))))))))))..........................((((.......))))..................... (-11.83 = -12.35 + 0.51)

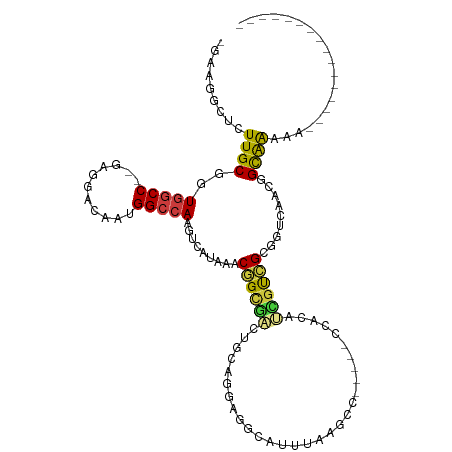

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:10:15 2011