
| Sequence ID | dm3.chrX |
|---|---|
| Location | 2,608,365 – 2,608,472 |
| Length | 107 |
| Max. P | 0.769916 |

| Location | 2,608,365 – 2,608,467 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 121 |
| Reading direction | forward |
| Mean pairwise identity | 54.11 |
| Shannon entropy | 0.71182 |
| G+C content | 0.56486 |
| Mean single sequence MFE | -32.78 |
| Consensus MFE | -13.14 |
| Energy contribution | -13.57 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.43 |
| Mean z-score | -0.93 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.600883 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 2608365 102 + 22422827 -------------------GCGCUGAUAAGGAUUUCAGACGAUUUCCUUACCUCCACGCCGGGAGAGGGCCAGAAGGACGUGCCCACCACGGAGAGCACAAAAUUGCAGAACCUGCACAGC -------------------..((((.((((((..((....))..)))))).(((....(((((...(((((........).)))).)).))).)))........(((((...))))))))) ( -30.90, z-score = -1.69, R) >droSec1.super_10 2369896 78 + 3036183 -------------------GCGUUGAUAUGCAUUUCAGACGAUUUCCGUACCUCCACGCCGGGAGAGGGUCAGGAGGUCGUGGAGAACCUGAACCGC------------------------ -------------------((((....)))).((((..(((((((((...((((..(....)..))))....))))))))).))))...........------------------------ ( -25.60, z-score = -0.73, R) >droYak2.chrX 16028972 120 - 21770863 ACAGCAAGCGACAGUGGCAGCGUC-AUAUGGAUUUCAAGCGAUUCCCUUGCCUCCUCACAGGGGGAGGCCCAGGGAGUCAAUGCUACCAAGGAGCUGGCAGACUUCGAGACUUUGUACUGC .((((........((((((((...-.............))((((((((.((((((((....))))))))..))))))))..))))))......))))((((((...........)).)))) ( -45.03, z-score = -1.65, R) >droEre2.scaffold_4690 34823 108 + 18748788 -------ACGACA-----AGCGAC-AUUUGGAUUUCAGAGGACUCUCUUGCCUCCUCGCAGGACGGGGCCCAAGGAGUCAUUGCUACCGAAGAGAGAGCAGACUUCGAGAACCCCUACAGG -------......-----.((((.-....(((....((((....))))....))))))).....((((..(..((((((..((((..(.....)..))))))))))..)..))))...... ( -29.60, z-score = 0.37, R) >consensus ___________________GCGUC_AUAUGGAUUUCAGACGAUUCCCUUACCUCCACGCAGGGAGAGGCCCAGGAAGUCAUGGCUACCAAGGAGAGCGCAGACUUCGAGAACCUGUACAGC ........................................(((((((..(((((.((....)).)))))...))))))).......................................... (-13.14 = -13.57 + 0.44)
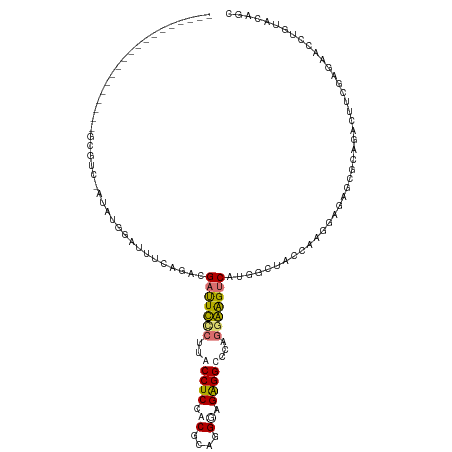


| Location | 2,608,370 – 2,608,472 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 53.98 |
| Shannon entropy | 0.71563 |
| G+C content | 0.56241 |
| Mean single sequence MFE | -34.08 |
| Consensus MFE | -13.14 |
| Energy contribution | -13.57 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.769916 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chrX 2608370 102 + 22422827 ------------------GAUAAGGAUUUCAGACGAUUUCCUUACCUCCACGCCGGGAGAGGGCCAGAAGGACGUGCCCACCACGGAGAGCACAAAAUUGCAGAACCUGCACAGCCGCAA ------------------..((((((..((....))..)))))).((((......))))..(((.........((((.(.(....).).)))).....(((((...)))))..))).... ( -28.50, z-score = -1.37, R) >droSec1.super_10 2369901 78 + 3036183 ------------------GAUAUGCAUUUCAGACGAUUUCCGUACCUCCACGCCGGGAGAGGGUCAGGAGGUCGUGGAGAACCUGAACCGCCGCAA------------------------ ------------------.....((..((((((((((((((...((((..(....)..))))....)))))))))(.....))))))..)).....------------------------ ( -23.80, z-score = -0.20, R) >droYak2.chrX 16028977 120 - 21770863 AAGCGACAGUGGCAGCGUCAUAUGGAUUUCAAGCGAUUCCCUUGCCUCCUCACAGGGGGAGGCCCAGGGAGUCAAUGCUACCAAGGAGCUGGCAGACUUCGAGACUUUGUACUGCCGCAA ..(((.(((((.(((.(((...(.((..((....((((((((.((((((((....))))))))..))))))))..(((((.(.....).)))))))..)).)))).)))))))).))).. ( -50.00, z-score = -2.72, R) >droEre2.scaffold_4690 34828 108 + 18748788 ------------AAGCGACAUUUGGAUUUCAGAGGACUCUCUUGCCUCCUCGCAGGACGGGGCCCAAGGAGUCAUUGCUACCGAAGAGAGAGCAGACUUCGAGAACCCCUACAGGCGCAA ------------..(((......((.((((..(((((((.((((...(((((.....)))))..))))))))).(((((..(.....)..))))).))..)))).))((....))))).. ( -34.00, z-score = -0.39, R) >consensus __________________CAUAUGGAUUUCAGACGAUUCCCUUACCUCCACGCAGGGAGAGGCCCAGGAAGUCAUGGCUACCAAGGAGAGCGCAAACUUCGAGAACCUGUACAGCCGCAA ..................................(((((((..(((((.((....)).)))))...)))))))............................................... (-13.14 = -13.57 + 0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:09:58 2011