
| Sequence ID | dm3.chrX |
|---|---|
| Location | 2,379,452 – 2,379,568 |
| Length | 116 |
| Max. P | 0.724009 |

| Location | 2,379,452 – 2,379,555 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 89.28 |
| Shannon entropy | 0.18547 |
| G+C content | 0.42999 |
| Mean single sequence MFE | -20.85 |
| Consensus MFE | -16.97 |
| Energy contribution | -16.89 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.724009 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 2379452 103 - 22422827 --CCAGUCUAUAACGUCAGCGACAACAAAGCCAACAAAUAUUGGCCCCGAAACGCGUUGAAGUAGCUUAUGACAUUGCCAUACUUAUAGACACUUAUAGCCACUA --...(((((((((.((((((........(((((......))))).........)))))).))....((((.......))))..))))))).............. ( -22.03, z-score = -2.38, R) >droEre2.scaffold_4644 2328789 99 - 2521924 CCAUAGUCUAUAACGUCAGCGACCACAAAGCCAACAAAUAUUGGCCCCGAAACGCGUUGAAGUAGCUUAAGACUUUGCC------AUAGAUACUCAUAGCCACUA .....((((((.((.((((((........(((((......))))).........)))))).)).((..........)).------)))))).............. ( -18.13, z-score = -1.28, R) >droYak2.chrX 15804463 97 + 21770863 --UCAGUCUAUAACGUCAGCGAUUACAAAGCCAACAAAUAUUGGCCCCGAAACGCGUUGGAGUAGCUUAUGACUUUGCC------AUAGAUACUCAUAGCAACUA --...(.((((.....(((((........(((((......))))).........)))))(((((..(((((.......)------)))).))))))))))..... ( -21.13, z-score = -1.31, R) >droSec1.super_10 2155317 96 - 3036183 --CCAGUCUAUAACGUCAGCGACAACAAAGCCAACAAAUAUUGG-CCCGAAUCGCGUUGAAGUAGCUUAUGACAUUGCC------AUAGAUACUUAUAGCCACUA --...(.(((((...((((((........(((((......))))-)........))))))((((..(((((.......)------)))).))))))))).).... ( -20.29, z-score = -1.76, R) >droSim1.chrX 1696225 98 - 17042790 UACCAGUCUAUAACGUCAGCGACAACAAAGCCAACAAAUAUUGG-CCCGUAUCGCGCUGAAGUAGCUUAUGACAUUGCC------AUAGAUACUUAUAGCCACUA .....(.(((((...((((((........(((((......))))-)........))))))((((..(((((.......)------)))).))))))))).).... ( -22.69, z-score = -2.15, R) >consensus __CCAGUCUAUAACGUCAGCGACAACAAAGCCAACAAAUAUUGGCCCCGAAACGCGUUGAAGUAGCUUAUGACAUUGCC______AUAGAUACUUAUAGCCACUA .....((((((.((.((((((........(((((......))))).........)))))).)).((..........)).......)))))).............. (-16.97 = -16.89 + -0.08)



| Location | 2,379,467 – 2,379,568 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 89.47 |
| Shannon entropy | 0.17498 |
| G+C content | 0.42684 |
| Mean single sequence MFE | -28.36 |
| Consensus MFE | -22.10 |
| Energy contribution | -22.90 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.552120 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 2379467 101 + 22422827 UCUAUAAGUAUGGCAAUGUCAUAAGCUACUUCAACGCGUUUCGGGGCCAAUAUUUGUUGGCUUUGUUGUCGCUGACGUUAUAGAC--UGGUAGCUA----AGAUACA (((.....((((((...))))))(((((((.....(((...((((((((((....))))))))))....)))....(((...)))--.))))))).----))).... ( -28.00, z-score = -1.41, R) >droEre2.scaffold_4644 2328804 97 + 2521924 ------UCUAUGGCAAAGUCUUAAGCUACUUCAACGCGUUUCGGGGCCAAUAUUUGUUGGCUUUGUGGUCGCUGACGUUAUAGACUAUGGUAGCUA----AGAUACA ------...........((((((.(((((......(((...((((((((((....))))))))))....)))...(((........))))))))))----))))... ( -28.80, z-score = -1.86, R) >droYak2.chrX 15804478 95 - 21770863 ------UCUAUGGCAAAGUCAUAAGCUACUCCAACGCGUUUCGGGGCCAAUAUUUGUUGGCUUUGUAAUCGCUGACGUUAUAGAC--UGAUAGCUA----AGAUACA ------((((((((..(((.....)))........(((...((((((((((....))))))))))....)))....)))))))).--.........----....... ( -24.50, z-score = -1.14, R) >droSec1.super_10 2155332 94 + 3036183 ------UCUAUGGCAAUGUCAUAAGCUACUUCAACGCGAUUCGGG-CCAAUAUUUGUUGGCUUUGUUGUCGCUGACGUUAUAGAC--UGGUAGCUA----AGAUACA ------..((((((...))))))(((((((.....(((((..(((-(((((....))))))))....)))))....(((...)))--.))))))).----....... ( -26.90, z-score = -1.72, R) >droSim1.chrX 1696240 98 + 17042790 ------UCUAUGGCAAUGUCAUAAGCUACUUCAGCGCGAUACGGG-CCAAUAUUUGUUGGCUUUGUUGUCGCUGACGUUAUAGAC--UGGUAGCUAGCUAAGAUACA ------..((((((...))))))((((((((((((((((((.(((-(((((....))))))))))))).)))))).(((...)))--.)))))))............ ( -33.60, z-score = -2.63, R) >consensus ______UCUAUGGCAAUGUCAUAAGCUACUUCAACGCGUUUCGGGGCCAAUAUUUGUUGGCUUUGUUGUCGCUGACGUUAUAGAC__UGGUAGCUA____AGAUACA ........((((((...))))))(((((((.....(((...((((((((((....))))))))))....)))...(......).....)))))))............ (-22.10 = -22.90 + 0.80)

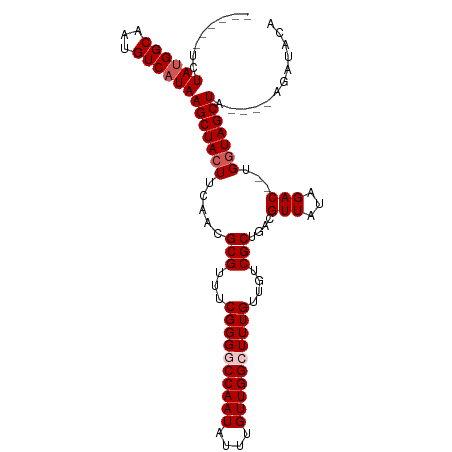

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:09:15 2011