| Sequence ID | dm3.chrX |
|---|---|
| Location | 2,290,413 – 2,290,511 |
| Length | 98 |
| Max. P | 0.938066 |

| Location | 2,290,413 – 2,290,511 |
|---|---|
| Length | 98 |
| Sequences | 8 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 54.59 |
| Shannon entropy | 0.89019 |
| G+C content | 0.53829 |
| Mean single sequence MFE | -16.59 |
| Consensus MFE | -7.15 |
| Energy contribution | -6.82 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.29 |
| Mean z-score | -0.86 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.938066 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 2290413 98 + 22422827 GCUACCUUGCUAUUUCGC-CUCCCCAUGCCAUUCGAUGC--CAUUCCACUGGCGCAGCUUAA--CGUCGAAAGUUGCUUUCG--UUACCAA--UACCUUACCUCACC- ((......))........-..............(((.((--((......))))(((((((..--......)))))))..)))--.......--..............- ( -15.80, z-score = -1.21, R) >droSec1.super_10 2070894 98 + 3036183 GCUACCUUGCUAUUCCGC-CUCCCCGUUCCAUUCGAUGC--CAUUCCACUGGCGCAGCUUAA--CGUCGAAAGUUGCUUUCG--UUACCUA--UACCUUACCUCACC- ((......))........-..............(((.((--((......))))(((((((..--......)))))))..)))--.......--..............- ( -15.80, z-score = -1.57, R) >droYak2.chrX 15725441 87 - 21770863 --------GAUCUUUUCC-CUUCCC---CUCUUUCAUUG--CAUUCCACUGGCGCAGCUUAA--CGUCGAAAGUUGCUUUCG--UUACCUA--UACCCCACCCCACC- --------..........-......---..........(--(.........))(((((((..--......))))))).....--.......--..............- ( -8.10, z-score = -0.44, R) >droAna3.scaffold_12929 2203058 100 + 3277472 ----GAGGGCCACUCGGCGCCACCCCUCUCACUUAAUUCUCCCCUUCAUUCUGGCAGCUUAG--CGCCGAAAGUUGCUUUCAGUUUACCCAGUUACGUUACCCAUU-- ----((((((.(((((((((((.............................)))).((...)--))))))..)).)))))).........................-- ( -20.35, z-score = -0.72, R) >dp4.chrXL_group1a 6118518 90 + 9151740 ------------CUUCCCCAGCACCUUCUCCCACCCUUGCCCUCCUGCCCGUUGCAGCUUAA--CGUUGAAAGUUGCUUUCA--CCCCCGG--CACGGCACUGGCACG ------------.........................((((..((((((.((.(((((((..--......)))))))....)--)....))--)).))....)))).. ( -18.50, z-score = -0.34, R) >droPer1.super_25 197134 89 - 1448063 ------------CUUCCCCUGCUCCUUCUCCCACCCUUGCCCUCCUGCCCGUUGCAGCUUAA--CGUUGAAAGUUGCUUUCA--CCCCCGG--CACGGCAC-GGCACC ------------.........................((((..((((((.((.(((((((..--......)))))))....)--)....))--)).))...-)))).. ( -19.20, z-score = -0.99, R) >droSim1.chrX 1627790 98 + 17042790 GCUACCUUGCUAUUCCGC-CUCCCCGUUCCAUUCGAUGC--CAUUCCACUGGCGCAGCUUAA--CGUCGAAAGUUGCUUUCG--UUACCUA--UACCUUACCUCACC- ((......))........-..............(((.((--((......))))(((((((..--......)))))))..)))--.......--..............- ( -15.80, z-score = -1.57, R) >droWil1.scaffold_180777 2001063 94 + 4753960 ----GUGUCCUUCUCACAUUGCAACGAAUUAUCGUCCUGUAUAGGGAAAACAUUCAGCUUAGUGCGCGGAAAGCUGG----AAGUUAAGUAACCCCCUCACC------ ----(((...(((((....((((((((....))))..))))..)))))(((.((((((((.(....)...)))))))----).)))............))).------ ( -19.20, z-score = -0.06, R) >consensus _______UGCUACUCCCC_CUCCCCGUCCCAUUCCAUUC__CAUUCCACUGGCGCAGCUUAA__CGUCGAAAGUUGCUUUCA__UUACCUA__UACCUCACCUCACC_ .....................................................(((((((..........)))))))............................... ( -7.15 = -6.82 + -0.33)
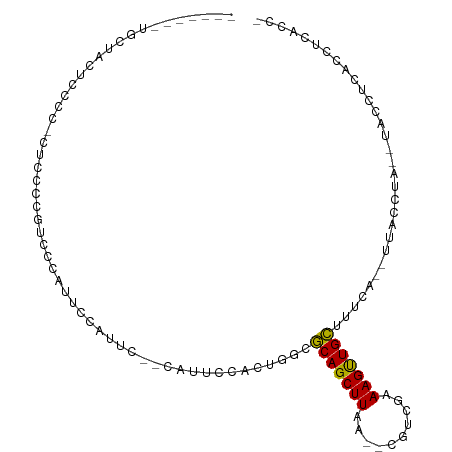


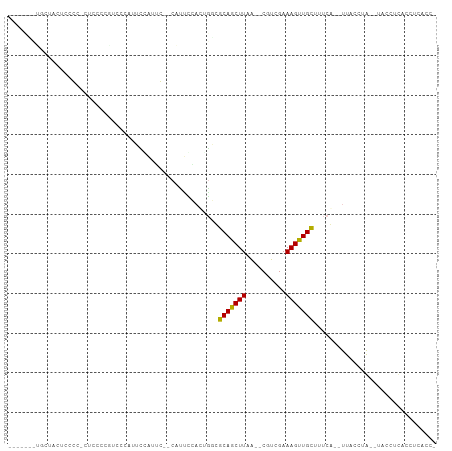
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:09:05 2011