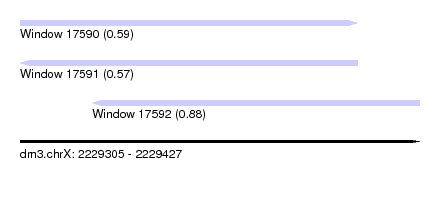

| Sequence ID | dm3.chrX |
|---|---|
| Location | 2,229,305 – 2,229,427 |
| Length | 122 |
| Max. P | 0.879772 |

| Location | 2,229,305 – 2,229,408 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 82.59 |
| Shannon entropy | 0.22936 |
| G+C content | 0.39177 |
| Mean single sequence MFE | -23.73 |
| Consensus MFE | -20.39 |
| Energy contribution | -20.17 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.86 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.589919 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
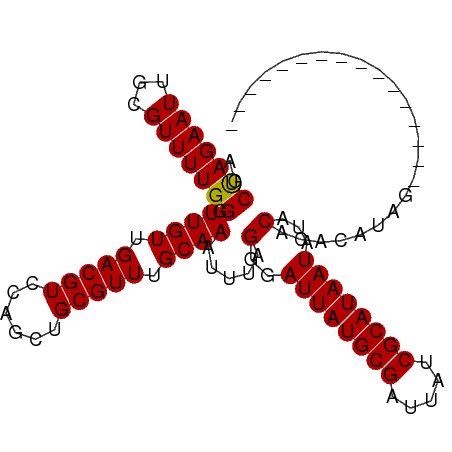
>dm3.chrX 2229305 103 + 22422827 AACCAGAAUUGCGUUUUGGGUUGUUGACGUCCAGCUGCGUUUGCAAAUUUAGGAAUUAUGCGAUUAUCGCAUAAUAAACAUACGUAUGUACAUAUGUAUAUAG ..(((((((...))))))).((((.(((((......))))).))))........((((((((.....))))))))....((((((((....)))))))).... ( -27.70, z-score = -1.92, R) >droSec1.super_10 2005633 87 + 3036183 AGCUAGAAUUGCGUUUUGGGUUGUUGACGUCCUGCUGCGUUUGCAAAUUUUGGGAUUAUGCGAUUAUCGCAUAAUAUACACACAUAG---------------- ..((((((((((((..((((.((....)).))))..)))).....)))))))).((((((((.....))))))))............---------------- ( -21.30, z-score = -0.75, R) >droSim1.chrX 1575492 87 + 17042790 AGCUAGAAUUGCGUUUUGGGUUGUUGACGUCCAGCUGCGUUUGCAAAUUUGGGGAUUAUGCGAUUAUCGCAUAAUAUACACACAUAG---------------- ..(((((.(((((....(.((.((((.....)))).)).).))))).)))))..((((((((.....))))))))............---------------- ( -22.20, z-score = -1.00, R) >consensus AGCUAGAAUUGCGUUUUGGGUUGUUGACGUCCAGCUGCGUUUGCAAAUUUAGGGAUUAUGCGAUUAUCGCAUAAUAUACACACAUAG________________ ..(((((((...))))))).((((.(((((......))))).)))).....(..((((((((.....))))))))...)........................ (-20.39 = -20.17 + -0.22)


| Location | 2,229,305 – 2,229,408 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 82.59 |
| Shannon entropy | 0.22936 |
| G+C content | 0.39177 |
| Mean single sequence MFE | -18.00 |
| Consensus MFE | -15.28 |
| Energy contribution | -15.40 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.567004 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
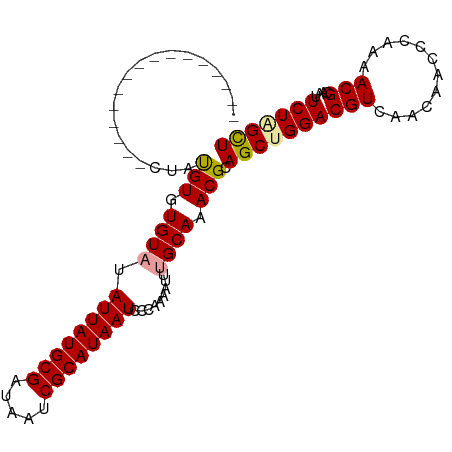
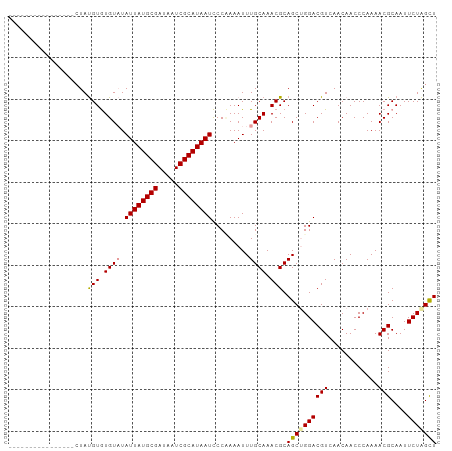
>dm3.chrX 2229305 103 - 22422827 CUAUAUACAUAUGUACAUACGUAUGUUUAUUAUGCGAUAAUCGCAUAAUUCCUAAAUUUGCAAACGCAGCUGGACGUCAACAACCCAAAACGCAAUUCUGGUU ......((((((((....))))))))..((((((((.....)))))))).((..(((((((....)))(((((...........)))....))))))..)).. ( -19.90, z-score = -1.18, R) >droSec1.super_10 2005633 87 - 3036183 ----------------CUAUGUGUGUAUAUUAUGCGAUAAUCGCAUAAUCCCAAAAUUUGCAAACGCAGCAGGACGUCAACAACCCAAAACGCAAUUCUAGCU ----------------...(((((....((((((((.....)))))))).......(((((.......)))))................)))))......... ( -16.10, z-score = -0.92, R) >droSim1.chrX 1575492 87 - 17042790 ----------------CUAUGUGUGUAUAUUAUGCGAUAAUCGCAUAAUCCCCAAAUUUGCAAACGCAGCUGGACGUCAACAACCCAAAACGCAAUUCUAGCU ----------------...(((((....((((((((.....))))))))..(((...((((....)))).)))................)))))......... ( -18.00, z-score = -1.62, R) >consensus ________________CUAUGUGUGUAUAUUAUGCGAUAAUCGCAUAAUCCCAAAAUUUGCAAACGCAGCUGGACGUCAACAACCCAAAACGCAAUUCUAGCU ...................(((.((((.((((((((.....)))))))).........)))).))).((((((((((............)))....))))))) (-15.28 = -15.40 + 0.12)

| Location | 2,229,327 – 2,229,427 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 57.88 |
| Shannon entropy | 0.68630 |
| G+C content | 0.38985 |
| Mean single sequence MFE | -17.90 |
| Consensus MFE | -6.25 |
| Energy contribution | -6.50 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.44 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.879772 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
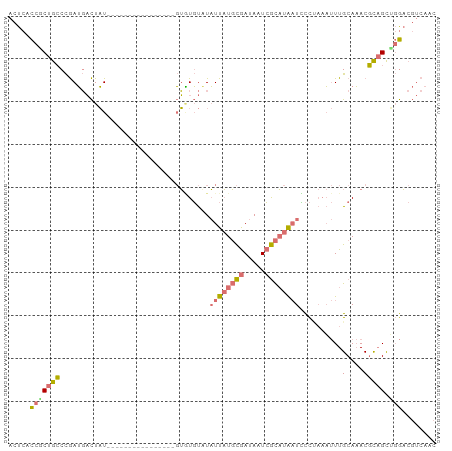
>dm3.chrX 2229327 100 - 22422827 ACUCACCACUGCCCGAUGACUAUAUACAUAUGUACAUACGUAUGUUUAUUAUGCGAUAAUCGCAUAAUUCCUAAAUUUGCAAACGCAGCUGGACGUCAAC .....(((((((.............((((((((....))))))))..((((((((.....))))))))................)))).)))........ ( -24.50, z-score = -3.11, R) >droSec1.super_10 2005655 84 - 3036183 ACUCACCGCUGCCCGAUGACUAU----------------GUGUGUAUAUUAUGCGAUAAUCGCAUAAUCCCAAAAUUUGCAAACGCAGCAGGACGUCAAC .....(((((((....((...((----------------...((...((((((((.....))))))))..))..))...))...))))).))........ ( -19.80, z-score = -1.37, R) >droSim1.chrX 1575514 84 - 17042790 ACUCACCGCUGCCCGAUGACUAU----------------GUGUGUAUAUUAUGCGAUAAUCGCAUAAUCCCCAAAUUUGCAAACGCAGCUGGACGUCAAC .....(((((((....((...((----------------...((...((((((((.....))))))))...)).))...))...))))).))........ ( -19.70, z-score = -1.49, R) >droGri2.scaffold_14853 4895075 91 - 10151454 ACUCAUCUCUAGCUAAGUUUUUUGUAUCAGAUUUUCUUUCAUUUCUUUUUUUUUAACAAUAAUUUGUUAUUU--CCUUGCACACUUUGUGUAU------- ....((((...((..........))...)))).....................((((((....))))))...--...(((((.....))))).------- ( -7.60, z-score = -0.87, R) >consensus ACUCACCGCUGCCCGAUGACUAU________________GUGUGUAUAUUAUGCGAUAAUCGCAUAAUCCCUAAAUUUGCAAACGCAGCUGGACGUCAAC .....(((((((...................................((((((((.....))))))))..........(....))))).)))........ ( -6.25 = -6.50 + 0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:08:58 2011