| Sequence ID | dm3.chrX |
|---|---|
| Location | 2,007,916 – 2,008,055 |
| Length | 139 |
| Max. P | 0.928349 |

| Location | 2,007,916 – 2,008,055 |
|---|---|
| Length | 139 |
| Sequences | 5 |
| Columns | 153 |
| Reading direction | reverse |
| Mean pairwise identity | 69.97 |
| Shannon entropy | 0.47716 |
| G+C content | 0.45626 |
| Mean single sequence MFE | -42.86 |
| Consensus MFE | -22.04 |
| Energy contribution | -22.96 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.37 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.928349 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
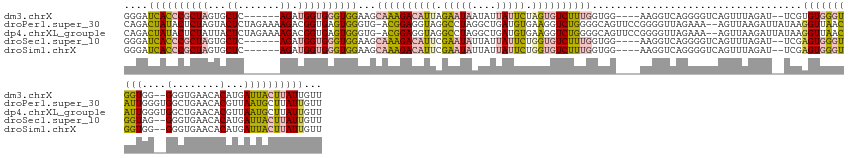
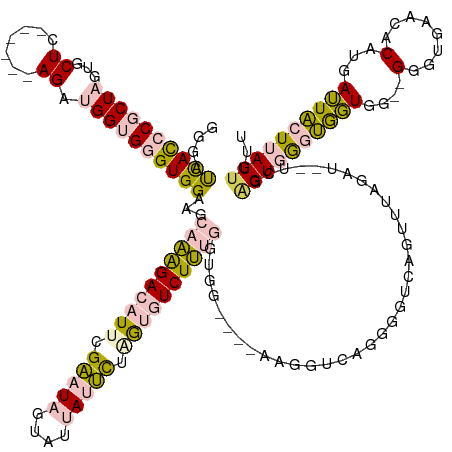
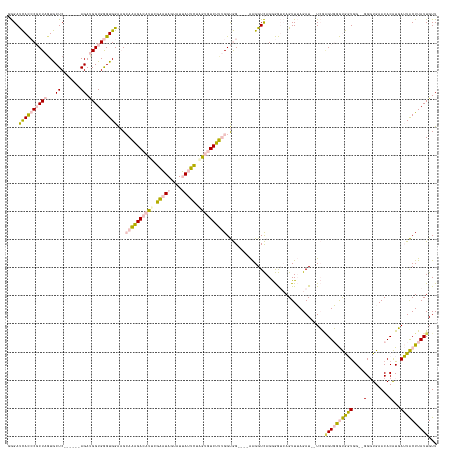
>dm3.chrX 2007916 139 - 22422827 GGGAUCACCCGCUAGUGCUC------AGAUGGUGGGUGGAAGCAAAGACAUUAGAAUAAUAUUAUUCUAGUGUCUUUGGUGG----AAGGUCAGGGGUCAGUUUAGAU--UCGUGUGGGUGGUGG--GGGUGAACACAUGAUUACUUAUUGUU ....((((((.(((.(((((------(.((..(((.(...(.((((((((((((((((....)))))))))))))))).)..----..).))).(((((......)))--)))).)))))).)))--)))))).................... ( -48.20, z-score = -4.20, R) >droPer1.super_30 218492 150 + 958394 CAGACUAUACUCUAGUACUCUAGAAAAGACGGUGAGUGGGUG-ACGGAGGUAGGCCUAGGCUGAUGUGAAGGUCUGGGGCAGUUCCGGGGUUAGAAA--AGUUAAGAUUAUAAGGUUAACAUUGGGUGGCUGAACACGUUAAUGCUUAUUGUU ..........(((((....)))))...(((((((((((..((-((((((.(.(.((((((((........)))))))).)).))))(...((((...--(.((((..(((......)))..)))).)..))))...))))).))))))))))) ( -40.80, z-score = -0.89, R) >dp4.chrXL_group1e 477212 150 - 12523060 CAGACUAUACUCUAUUACUCUAGAAAAGACGGUGAGUGGGUG-ACGGAGGUAGGCCUAGGCUGAUGUGAAGGUCUGGGGCAGUUCCGGGGUUAGAAA--AGUUAAGAUUAUAAGGUUAACAUUGGGUGGCUGAACACGUUAAUGCUUAUUGUU ..........((((......))))...(((((((((((..((-((((((.(.(.((((((((........)))))))).)).))))(...((((...--(.((((..(((......)))..)))).)..))))...))))).))))))))))) ( -37.70, z-score = -0.10, R) >droSec1.super_10 1792353 139 - 3036183 GGGAUCACCCGCUAGUGCUC------AGAUGGUGGGUGGAAGCAAAGACAUUCGAAUAUUAUUAUUCUGGUGUCUUUGGUGG----AAGGUCAGGGGUCAGUUUAGAU--UCGAGUGGGUGGUAG--GGGUGAACACAUGAUUACUUAUUGUU ..((((((((((((......------...)))))))).....(((((((((..(((((....)))))..)))))))))....----..))))..(((((......)))--)).((((((((((..--.((....).)...))))))))))... ( -43.50, z-score = -2.39, R) >droSim1.chrX 1396616 139 - 17042790 GGGAUCACCCGCUAGUGCUC------AGAUGGUGGGUGGAAGCAAAGACAUUCGAAUAUUAUUAUUCUGGUGUCUUUGGUGG----AAGGUCAGGGGUCAGUUUAGAU--UCGAGUGGGUGGUGG--GGGUGAACACAUGAUUACUUAUUGUU ..((((((((((((......------...)))))))).....(((((((((..(((((....)))))..)))))))))....----..))))..(((((......)))--)).((((((((((.(--..(....)...).))))))))))... ( -44.10, z-score = -2.42, R) >consensus GGGAUCACCCGCUAGUGCUC______AGAUGGUGGGUGGAAGCAAAGACAUUCGAAUAGUAUUAUUCUAGUGUCUUUGGUGG____AAGGUCAGGGGUCAGUUUAGAU__UCGAGUGGGUGGUGG__GGGUGAACACAUGAUUACUUAUUGUU ....((((((((((...((.......)).))))))))))...((((((((((.(((((....))))).))))))))))...................................((((((((((.....(.....).....))))))))))... (-22.04 = -22.96 + 0.92)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:08:18 2011