
| Sequence ID | dm3.chrX |
|---|---|
| Location | 1,992,161 – 1,992,213 |
| Length | 52 |
| Max. P | 0.932457 |

| Location | 1,992,161 – 1,992,213 |
|---|---|
| Length | 52 |
| Sequences | 6 |
| Columns | 54 |
| Reading direction | forward |
| Mean pairwise identity | 74.68 |
| Shannon entropy | 0.48299 |
| G+C content | 0.64613 |
| Mean single sequence MFE | -11.97 |
| Consensus MFE | -7.40 |
| Energy contribution | -6.76 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.57 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.909792 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
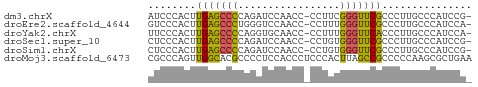
>dm3.chrX 1992161 52 + 22422827 AUCCCACUUGAGCCCCAGAUCCAACC-CCUUCGGGUUCGCCCUUGCCCAUCCG- .........((((((.((........-.))..))))))...............- ( -8.30, z-score = -0.86, R) >droEre2.scaffold_4644 1965103 52 + 2521924 GUCCCACUUGAGCCCUGGGUCCAACC-CCUUUGGGUUCGCCCUUGCCCAUCCA- .........((((((.(((......)-))...))))))...............- ( -14.70, z-score = -1.23, R) >droYak2.chrX 15438143 52 - 21770863 UUCCCACUUGAGCCCCAGGUGCAACC-CCUUUGGGUUCACCCUUGCCCAUCCA- ........((.((....((((.((((-(....)))))))))...)).))....- ( -13.90, z-score = -1.73, R) >droSec1.super_10 1776650 52 + 3036183 CUCCCACUUGAGCCCCAGAUCCAACC-CCUGUGGGUUCGCCCUUGCCCAUCCG- .........(((((((((........-.))).))))))...............- ( -12.60, z-score = -2.25, R) >droSim1.chrX 1400933 52 + 17042790 CUCCCACUUGAGCCCCAGAUCCAACC-CCUGUGGGUUCGCCCUUGCCCAUCCG- .........(((((((((........-.))).))))))...............- ( -12.60, z-score = -2.25, R) >droMoj3.scaffold_6473 16762959 54 - 16943266 CGCCCAGUUGGCACGCCCCUCCACCCUCCCACUUAGCCGCCCCCAAGCGCUGAA .(((.....)))....................(((((.((......))))))). ( -9.70, z-score = -0.59, R) >consensus CUCCCACUUGAGCCCCAGAUCCAACC_CCUUUGGGUUCGCCCUUGCCCAUCCA_ ........(((((((.................)))))))............... ( -7.40 = -6.76 + -0.64)


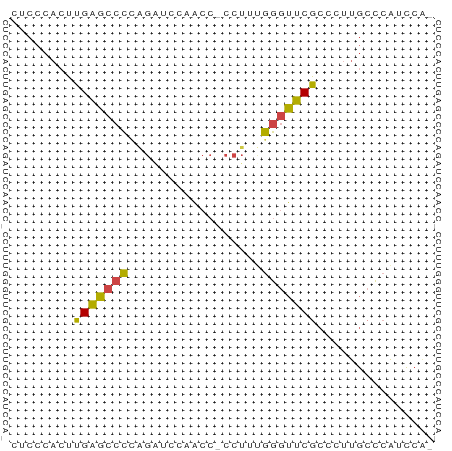
| Location | 1,992,161 – 1,992,213 |
|---|---|
| Length | 52 |
| Sequences | 6 |
| Columns | 54 |
| Reading direction | reverse |
| Mean pairwise identity | 74.68 |
| Shannon entropy | 0.48299 |
| G+C content | 0.64613 |
| Mean single sequence MFE | -19.38 |
| Consensus MFE | -9.67 |
| Energy contribution | -9.90 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.46 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.932457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 1992161 52 - 22422827 -CGGAUGGGCAAGGGCGAACCCGAAGG-GGUUGGAUCUGGGGCUCAAGUGGGAU -....(((((.((..(.(((((....)-)))).)..))...)))))........ ( -20.70, z-score = -3.54, R) >droEre2.scaffold_4644 1965103 52 - 2521924 -UGGAUGGGCAAGGGCGAACCCAAAGG-GGUUGGACCCAGGGCUCAAGUGGGAC -....(((((..((((.(((((....)-)))).).)))...)))))........ ( -21.20, z-score = -2.40, R) >droYak2.chrX 15438143 52 + 21770863 -UGGAUGGGCAAGGGUGAACCCAAAGG-GGUUGCACCUGGGGCUCAAGUGGGAA -....(((((..((((((((((....)-)))).)))))...)))))........ ( -22.70, z-score = -3.13, R) >droSec1.super_10 1776650 52 - 3036183 -CGGAUGGGCAAGGGCGAACCCACAGG-GGUUGGAUCUGGGGCUCAAGUGGGAG -....(((((.((..(.(((((....)-)))).)..))...)))))........ ( -18.00, z-score = -1.84, R) >droSim1.chrX 1400933 52 - 17042790 -CGGAUGGGCAAGGGCGAACCCACAGG-GGUUGGAUCUGGGGCUCAAGUGGGAG -....(((((.((..(.(((((....)-)))).)..))...)))))........ ( -18.00, z-score = -1.84, R) >droMoj3.scaffold_6473 16762959 54 + 16943266 UUCAGCGCUUGGGGGCGGCUAAGUGGGAGGGUGGAGGGGCGUGCCAACUGGGCG .....((((..(.(((((((...(.(.....).)...))).))))..)..)))) ( -15.70, z-score = 0.58, R) >consensus _CGGAUGGGCAAGGGCGAACCCAAAGG_GGUUGGAUCUGGGGCUCAAGUGGGAG .....(((((..((((.((((.......)))).).)))...)))))........ ( -9.67 = -9.90 + 0.23)


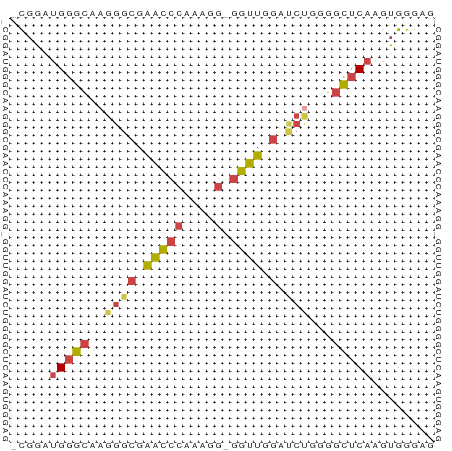
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:08:12 2011