
| Sequence ID | dm3.chrX |
|---|---|
| Location | 1,984,516 – 1,984,636 |
| Length | 120 |
| Max. P | 0.618156 |

| Location | 1,984,516 – 1,984,636 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 132 |
| Reading direction | forward |
| Mean pairwise identity | 71.45 |
| Shannon entropy | 0.52325 |
| G+C content | 0.53627 |
| Mean single sequence MFE | -40.22 |
| Consensus MFE | -22.19 |
| Energy contribution | -21.98 |
| Covariance contribution | -0.21 |
| Combinations/Pair | 1.52 |
| Mean z-score | -1.06 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.618156 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
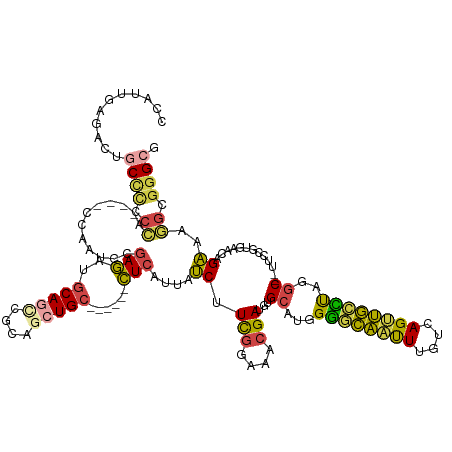
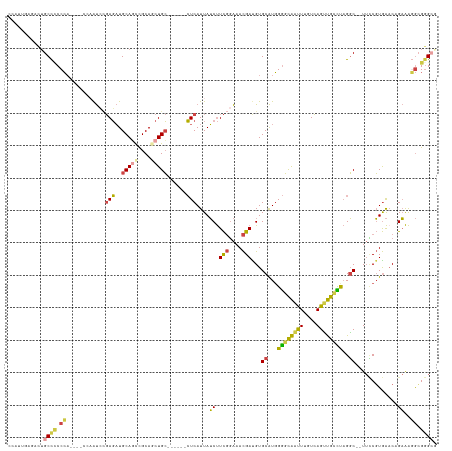
>dm3.chrX 1984516 120 + 22422827 CCAUUGAGACUGCCCCCCA----CCAAUCCGAGAUGCAGCCGCAGCUGC------CUCAUUAUCUUCGGAAACGAUGUGCAUGGGGCAAUUUGUCAGUUGCUUAGGC--UUCCGUGAACAGAAACGCGGGCA ..(((((.(.((((((.((----(......(((..(((((....)))))------))).......(((....))).)))...))))))...).))))).((....))--.((((((........)))))).. ( -38.50, z-score = -0.52, R) >droAna3.scaffold_13117 5135617 125 + 5790199 CAACUGCCACUGCUUCCGC-------ACAUCAUCUGCAUCUGCAUCUGAAUCUGAAUCAUUAGCCUUUCGAAUGAUGUGCGUUGAUUGCCUAAUGAGGCAAUUUGUCAGUAGCUUAGACCGCGGCAAGGGCC ....(((((((((...(((-------(((((((((((....)))...(((.((((....))))...)))).))))))))))..((((((((....)))))))).).)))).((.......))))))...... ( -38.10, z-score = -1.23, R) >droEre2.scaffold_4644 1957276 116 + 2521924 CCAUUGAAUCUGCCGCCCA-------AUCAGAGAUGCAGCUGCAGCUGC------CUCAUUAUCUUCGGAAACGACGUGCAUGGGGCAAUUUGUCAGUUGCCUAGGC--UUCCGUGA-CUGAAAGGGGGGAG ............((.(((.-------.((((....((((((((((.(((------(((((...(.(((....))).)...))))))))..))).)))))))...((.--..))....-))))..))).)).. ( -43.30, z-score = -1.53, R) >droYak2.chrX 15430291 130 - 21770863 CCCUGGAAAUGGCCCCCCAAAUCCAAGUACGAGAUGCAACCGCAACUGCAACUGCCUCAUUAUCUUCGGAAACGACGUGCAUGGGGCAAUUUGUCAGCUGGCCAGGC--UUCCGUGAACCGAAAGGCGGGAG (((.((((.(((((....................(((....))).((((((.((((((((...(.(((....))).)...))))))))..))).)))..)))))...--)))).....((....)).))).. ( -45.10, z-score = -1.40, R) >droSec1.super_10 1769056 120 + 3036183 CCUUUAAGACUGCCCCCCA----CCAAUCCGAGAUGCAGUCGCAGCUGC------CUCAUUAUCUUCGGAAACGAUGUGCAUGGGGCAAUUUGUCAGUUGCCUAGGC--UUUCGUGAACAGAAAGGCGAGCA .......(((((((((.((----(......(((..(((((....)))))------))).......(((....))).)))...))))))....)))...(((...(.(--(((((....).))))).)..))) ( -37.70, z-score = -0.99, R) >droSim1.chrX 1364492 120 + 17042790 CCUUUAAGACUGCCCCCCA----CCAAUCCGAGAUGCAGUCGCAGCUGC------CUCAUUAUCUUCGGAAACGAUGUGCAUGGGGCAAUUUGUCAGUUGCCUAGGC--UUCCGUGAACAGAAAGGCGGGCG ...........((((.((.----.......(((..(((((....)))))------)))....((.(((....)))(((.(((((((((((......)))))).....--..))))).)))))..)).)))). ( -38.61, z-score = -0.67, R) >consensus CCAUUGAGACUGCCCCCCA____CCAAUCCGAGAUGCAGCCGCAGCUGC______CUCAUUAUCUUCGGAAACGAUGUGCAUGGGGCAAUUUGUCAGUUGCCUAGGC__UUCCGUGAACAGAAAGGCGGGCG ...........((((.((............(((..(((((....)))))......)))....((.(((....)))...((...((((((((....))))))))..)).............))..)).)))). (-22.19 = -21.98 + -0.21)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:08:11 2011