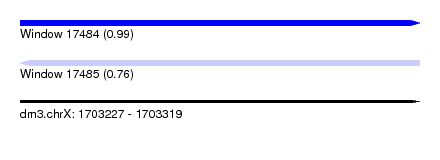

| Sequence ID | dm3.chrX |
|---|---|
| Location | 1,703,227 – 1,703,319 |
| Length | 92 |
| Max. P | 0.986252 |

| Location | 1,703,227 – 1,703,319 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 65.54 |
| Shannon entropy | 0.61904 |
| G+C content | 0.56180 |
| Mean single sequence MFE | -32.36 |
| Consensus MFE | -18.08 |
| Energy contribution | -16.56 |
| Covariance contribution | -1.52 |
| Combinations/Pair | 1.62 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.23 |
| SVM RNA-class probability | 0.986252 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
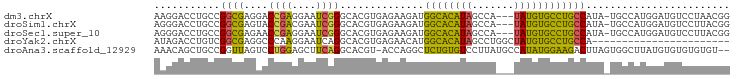
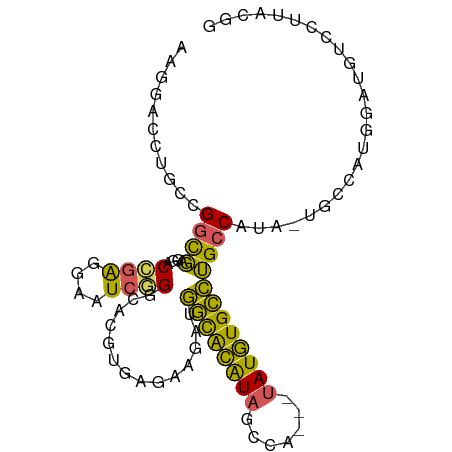
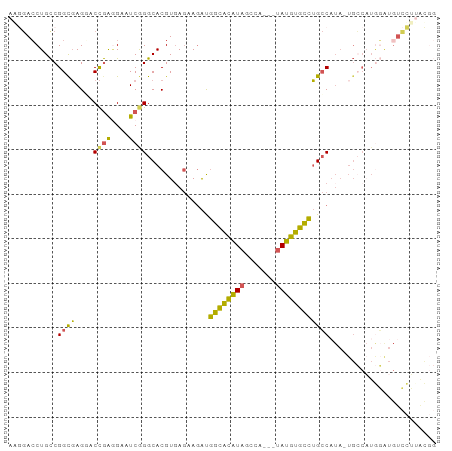
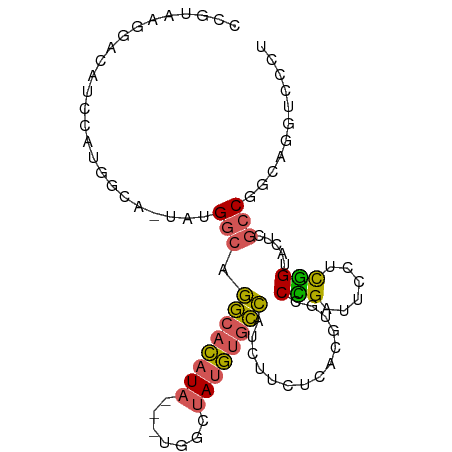
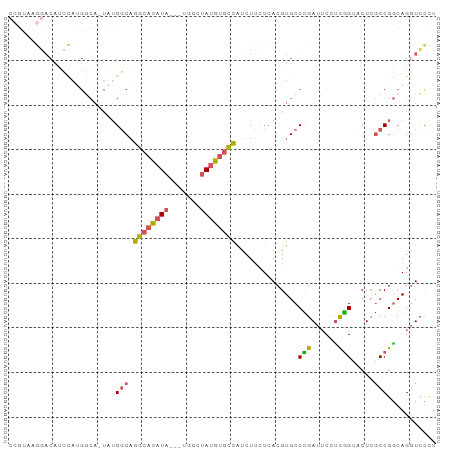
>dm3.chrX 1703227 92 + 22422827 AAGGACCUGCCGGCGAGGACCGAGGAAUCGGGCACGUGAGAAGAUGGCACAUAGCCA---UAUGUGCCUGCCAUA-UGCCAUGGAUGUCCUAACGG .(((((...((((((.....(((....)))(((((....).....((((((((....---))))))))))))...-))))..))..)))))..... ( -35.20, z-score = -2.17, R) >droSim1.chrX 1196667 92 + 17042790 AGGGACCUGCCGGCGAGUACCGACGAAUCGGGCACGUGAGAAGAUGGCACAUAGCCA---UAUGUGCCUGCCAUA-UGCCAUGGAUGUCCUUACGG ((((((...((((((.....(((....)))(((((....).....((((((((....---))))))))))))...-))))..))..)))))).... ( -34.50, z-score = -1.76, R) >droSec1.super_10 1498314 92 + 3036183 AGGGACCUGCCGGCGAGAACCGAGGAAUCGGGCACGUGAGAAGAUGGCACAUAGCCA---UAUGUGCCUGCCAUA-UGCCAUGGAUGUCCUUACGG ((((((...((((((.....(((....)))(((((....).....((((((((....---))))))))))))...-))))..))..)))))).... ( -35.50, z-score = -2.17, R) >droYak2.chrX 1548894 73 + 21770863 AUAGACCUGUCGGCGAGGCCCAAGGAAUCAGGCACGUGAGAACAUGGCACAUAGCCUGGCUAUGUGCCUGCCA----------------------- .....(((...(((...)))..))).....(((((....).....((((((((((...)))))))))))))).----------------------- ( -30.00, z-score = -2.26, R) >droAna3.scaffold_12929 1721308 93 - 3277472 AAACAGCUGCCGGUUAGUCCUGGAGCUUCAGGCACGU-ACCAGGCUCUGUGUCCUUAUGCCAUAUGGAAGACUUAGUGGCUUAUGUGUGUGUGU-- ....((((.((((......)))))))).((.((((((-(...(((((((.(((......((....))..))).))).))))))))))).))...-- ( -26.60, z-score = 0.46, R) >consensus AAGGACCUGCCGGCGAGGACCGAGGAAUCGGGCACGUGAGAAGAUGGCACAUAGCCA___UAUGUGCCUGCCAUA_UGCCAUGGAUGUCCUUACGG ...........((((....((((....))))..............((((((((.......))))))))))))........................ (-18.08 = -16.56 + -1.52)
| Location | 1,703,227 – 1,703,319 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 65.54 |
| Shannon entropy | 0.61904 |
| G+C content | 0.56180 |
| Mean single sequence MFE | -28.96 |
| Consensus MFE | -15.18 |
| Energy contribution | -14.86 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.43 |
| Mean z-score | -0.90 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.764989 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
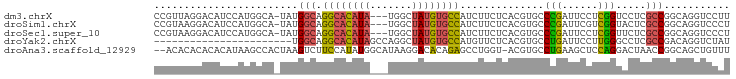
>dm3.chrX 1703227 92 - 22422827 CCGUUAGGACAUCCAUGGCA-UAUGGCAGGCACAUA---UGGCUAUGUGCCAUCUUCUCACGUGCCCGAUUCCUCGGUCCUCGCCGGCAGGUCCUU .....(((((....((((((-((((((.........---..)))))))))))).........((((.((....))(((....))))))).))))). ( -30.80, z-score = -0.88, R) >droSim1.chrX 1196667 92 - 17042790 CCGUAAGGACAUCCAUGGCA-UAUGGCAGGCACAUA---UGGCUAUGUGCCAUCUUCUCACGUGCCCGAUUCGUCGGUACUCGCCGGCAGGUCCCU ((....))........((((-..(((..((((((((---....))))))))......)))..)))).(((..((((((....))))))..)))... ( -33.50, z-score = -1.42, R) >droSec1.super_10 1498314 92 - 3036183 CCGUAAGGACAUCCAUGGCA-UAUGGCAGGCACAUA---UGGCUAUGUGCCAUCUUCUCACGUGCCCGAUUCCUCGGUUCUCGCCGGCAGGUCCCU ......((((....((((((-((((((.........---..)))))))))))).........((((.((....))(((....))))))).)))).. ( -29.90, z-score = -0.58, R) >droYak2.chrX 1548894 73 - 21770863 -----------------------UGGCAGGCACAUAGCCAGGCUAUGUGCCAUGUUCUCACGUGCCUGAUUCCUUGGGCCUCGCCGACAGGUCUAU -----------------------.(((.((((((((((...))))))))))((((....)))))))........(((((((.......))))))). ( -29.20, z-score = -1.82, R) >droAna3.scaffold_12929 1721308 93 + 3277472 --ACACACACACAUAAGCCACUAAGUCUUCCAUAUGGCAUAAGGACACAGAGCCUGGU-ACGUGCCUGAAGCUCCAGGACUAACCGGCAGCUGUUU --.............(((......(((((((....))....))))).....(((.(((-..((.((((......))))))..)))))).))).... ( -21.40, z-score = 0.20, R) >consensus CCGUAAGGACAUCCAUGGCA_UAUGGCAGGCACAUA___UGGCUAUGUGCCAUCUUCUCACGUGCCCGAUUCCUCGGUACUCGCCGGCAGGUCCCU ........................(((.((((((((.......))))))))..............(((......))).....)))........... (-15.18 = -14.86 + -0.32)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:07:28 2011