
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 845,827 – 845,916 |
| Length | 89 |
| Max. P | 0.545255 |

| Location | 845,827 – 845,916 |
|---|---|
| Length | 89 |
| Sequences | 8 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 67.84 |
| Shannon entropy | 0.64147 |
| G+C content | 0.44395 |
| Mean single sequence MFE | -11.60 |
| Consensus MFE | -5.50 |
| Energy contribution | -5.51 |
| Covariance contribution | 0.02 |
| Combinations/Pair | 1.11 |
| Mean z-score | -0.78 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.545255 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
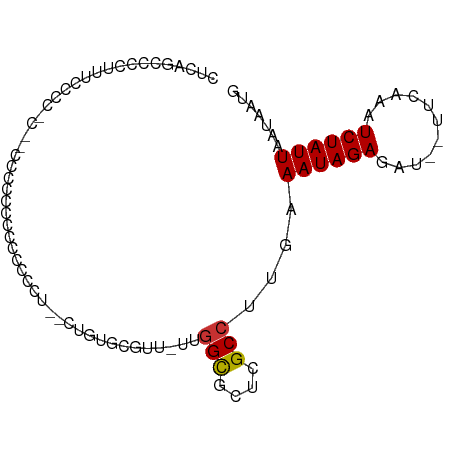
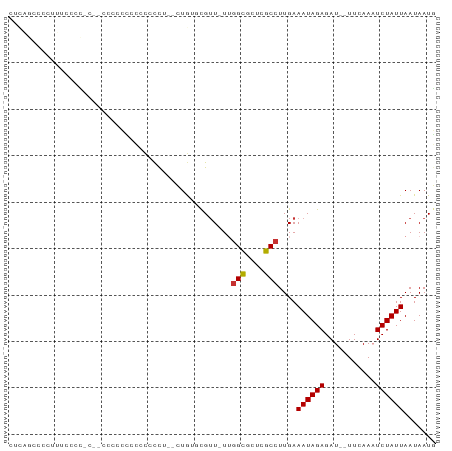
>dm3.chr2L 845827 89 - 23011544 CUCAGCGCCUUCCCCCUCCACACCCCCCACCCCUUCCUGCGCGUU-UUGGCGCUCGCCUUGAAAUAGAGAU--UUGAAAUCUAUUAAUAAUG .(((((((..............................))))...-..(((....))).)))((((((...--......))))))....... ( -12.31, z-score = -0.91, R) >droPer1.super_1 7129687 85 - 10282868 -------GUUUUUGUAUCGCCCCCCACUCCCCUUUUGUGUUUGUUUUGGGCUCUCGCCUUGAAAUAGAGAUUUUUCAAAUCUAUUAAUAAUU -------...((((.(((......(((.........)))((((((((((((....))).))))))))))))....))))............. ( -11.60, z-score = -0.71, R) >dp4.chr4_group3 5638725 85 - 11692001 -------GUUUUUGUAUCGCCCCCCACUCCCCUUUUGUGUUUGUUUUGGGCUCUCGCCUUGAAAUAGAGAUUUUUCAAAUCUAUUAAUAAUU -------...((((.(((......(((.........)))((((((((((((....))).))))))))))))....))))............. ( -11.60, z-score = -0.71, R) >droAna3.scaffold_12916 2735878 82 - 16180835 CUUGGCCAACUGCCCU----CCCCCACCACCCC---CUAGCUGUU-UUGGCGCUCGCCUUGAAAUAGAGAU--UUCAAUUCUAUUAAUAAUG ...(((.....)))..----.............---....(((((-(((((....)))..)))))))(((.--......))).......... ( -11.70, z-score = -0.38, R) >droEre2.scaffold_4929 902588 85 - 26641161 CUCAGCCCCCUUGGCCCC--CUCCCCUCCCCCCG--CUGAGCGUU-UUGGCGCUUGCCUUGAAAUAGAGAU--UUGAAAUCUAUUAAUAAUG .((((((.....)))...--.............(--(.((((((.-...))))))))..)))((((((...--......))))))....... ( -15.50, z-score = -1.19, R) >droYak2.chr2L 829536 83 - 22324452 CUCAGCCCCAUUCCCU----CCCCUCUCCUCCCU--CUGCGCGAU-UUGGCGCUUGCCUUGAAAUAGAGAU--UUGAAAUCUAUUAAUAAUG ................----..............--..((((...-...)))).........((((((...--......))))))....... ( -9.60, z-score = 0.26, R) >droSec1.super_14 816468 86 - 2068291 CUCAGCCCCUUUCCCCCUC-CAUCCCCCACCCCA--CUGAGCGUU-UUGGCGCUCGCCUUGAAAUAGAGAU--UUGAAAUCUAUUAAUAAUG ...................-..............--..((((((.-...)))))).......((((((...--......))))))....... ( -12.50, z-score = -1.30, R) >droWil1.scaffold_180772 6610677 70 + 8906247 --------------------GUCUCUCUUUCUUUGUGUGUAUGUUAUUUGUUUUUGCCUUGAAAUAGAGAU--UUCAAUUCUAUUAAUAAUG --------------------((((((.((((...(((...(((.....)))...)))...)))).))))))--................... ( -8.00, z-score = -1.30, R) >consensus CUCAGCCCCUUUCCCC_C__CCCCCCCCCCCCCU__CUGUGCGUU_UUGGCGCUCGCCUUGAAAUAGAGAU__UUCAAAUCUAUUAAUAAUG ................................................(((....)))....((((((...........))))))....... ( -5.50 = -5.51 + 0.02)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:07:11 2011