
| Sequence ID | dm3.chrX |
|---|---|
| Location | 1,641,454 – 1,641,556 |
| Length | 102 |
| Max. P | 0.612015 |

| Location | 1,641,454 – 1,641,556 |
|---|---|
| Length | 102 |
| Sequences | 8 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 80.03 |
| Shannon entropy | 0.37020 |
| G+C content | 0.47870 |
| Mean single sequence MFE | -32.29 |
| Consensus MFE | -12.56 |
| Energy contribution | -12.56 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.612015 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 1641454 102 + 22422827 UGCCACCGUUUGUGUACAUUCUGACAAAUGUGUAUAAAUAUUUUCGCUGCCCGCGU----GAGCUCACUGAGCUGAAUACCGACACCUAUACAG-GUGGCAUCGGUG- ((((((((((((((((((((......))))))))))))).....(((.....)))(----.(((((...))))).).................)-))))))......- ( -28.80, z-score = -1.27, R) >droSim1.chrX 1150588 97 + 17042790 UGCCACCGUUUGUGUACAUUCUGACAAAUGUGUAUAAAUAUUUUCGCUGCCCGCGU----GAGCUCAGUGAGCUGAAUACCGACACCUAUACAG-GUGGCAU------ ((((((((((((((((((((......)))))))))))))...(((((((...((..----..)).))))))).....................)-)))))).------ ( -33.30, z-score = -3.46, R) >droSec1.super_10 1436427 97 + 3036183 UGCCACCGUUUGUGUACAUUCUGACAAAUGUGUAUAAAUAUUUUCGCUGCCCGCGU----GAGCUCAGUGAGCUGAAUACCGACACCUAUACAG-GUGGCAU------ ((((((((((((((((((((......)))))))))))))...(((((((...((..----..)).))))))).....................)-)))))).------ ( -33.30, z-score = -3.46, R) >droYak2.chrX 1484426 103 + 21770863 UGCCACCGUUUGUGUACAUUCUGACAAAUGUGUAUAAAUAUUUUCGCUCCCCGCGU----GAGCUCAGUGAGCUGAAUACCGACACCUAUACGA-GUACAGGUGGCAU ((((((((((((((((((((......)))))))))))))......((((...((..----..))((((....))))................))-))...))))))). ( -33.80, z-score = -3.34, R) >droEre2.scaffold_4644 1619087 97 + 2521924 UGCCACCGUUUGUGUACAUUCUGACAAAUGUGUAUAAAUAUUUUCGCUCCCCGCGU----GAGCUCAGUGAGCUGAAUGGCGACACCUAUACAG-GUGGCAU------ ((((((((((((((((((((......)))))))))))))....(((((....((..----..))((((....))))..)))))..........)-)))))).------ ( -35.50, z-score = -3.71, R) >droAna3.scaffold_12929 1517090 106 + 3277472 UGCCAACGUUUGUGGACAUUCAGACAAAUGUGUAUAAAUAUUUUCGUUCCCCAAUCCACCGAGCUCACUGAGCUGAACACCGACACCUACCGAGAGCAGGGUGUGU-- .......(((((((((......(((((((((......))))))..)))......)))))..(((((...)))))))))....((((((..(....)..))))))..-- ( -23.90, z-score = -0.11, R) >dp4.chrXL_group1a 4813673 99 - 9151740 UGCCAACGUUUGUAUACAUUCAUACAAAUGUGUAUAAAUAUUUUCGCUACCAACUCC---GCGCUCCGUGGGCUGCAUGGUGACACCUUGCGGAGGGGGAGG------ .((.((..((((((((((((......))))))))))))..))...))......((((---.(.(((((..((.....((....)).))..))))).))))).------ ( -35.20, z-score = -2.58, R) >droPer1.super_11 1930177 99 - 2846995 UGCCAACGUUUGUAUACAUUCAUACAAAUGUGUAUAAAUAUUUUCGCUACCAACUCC---GCGCUCCGUGGGCUGCAUGGCGACACCUUGCGGAGGGGGAGG------ .((.((..((((((((((((......))))))))))))..))...))......((((---.(.(((((..((.((........)).))..))))).))))).------ ( -34.50, z-score = -2.30, R) >consensus UGCCACCGUUUGUGUACAUUCUGACAAAUGUGUAUAAAUAUUUUCGCUCCCCGCGU____GAGCUCAGUGAGCUGAAUACCGACACCUAUACAG_GUGGCAU______ .......(((((((((((((......)))))))))))))......................(((((...))))).................................. (-12.56 = -12.56 + 0.00)

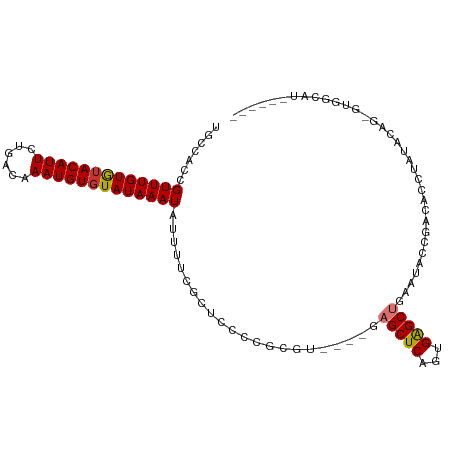

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:07:12 2011