| Sequence ID | dm3.chrX |
|---|---|
| Location | 1,563,544 – 1,563,640 |
| Length | 96 |
| Max. P | 0.942790 |

| Location | 1,563,544 – 1,563,640 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 62.55 |
| Shannon entropy | 0.70004 |
| G+C content | 0.49728 |
| Mean single sequence MFE | -36.44 |
| Consensus MFE | -12.95 |
| Energy contribution | -9.29 |
| Covariance contribution | -3.66 |
| Combinations/Pair | 1.87 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.48 |
| SVM RNA-class probability | 0.942790 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
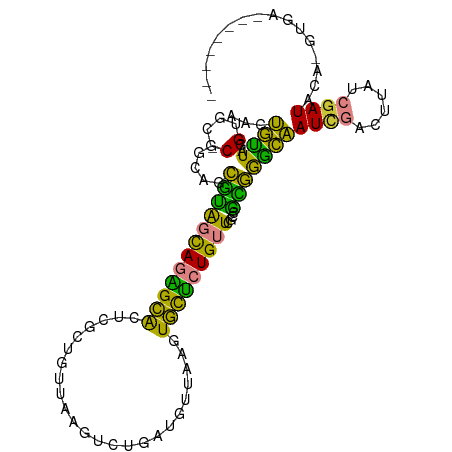
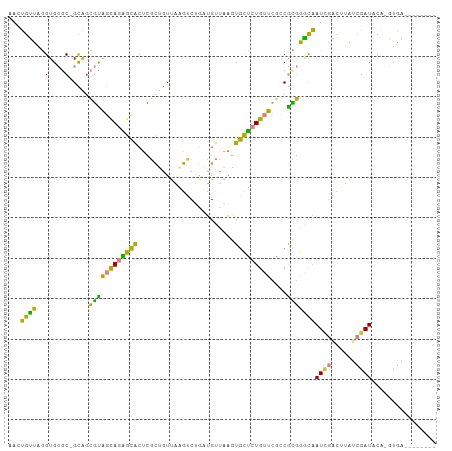
>dm3.chrX 1563544 96 - 22422827 AAAUGUUAAGUGCGC-GCAGCGCAGCAGAGCACUCGCUGUUAAGUCUGAUGUUAAAUGCUCUGUUCGCCGCGGGCAAUCGACUUAUCGAUACA-GUGA-------- ...(((....(((.(-((.(((.(((((((((...((.((((....))))))....)))))))))))).))).)))(((((....))))))))-....-------- ( -32.80, z-score = -0.95, R) >droEre2.scaffold_4644 1548742 99 - 2521924 CACUGUUAGGUGCGC-GCA-CGUAGCAGAGCAGUCGCUGUUAAGUC---UGUUAAGUGCUCUGUUCGCCGUGGACAAUCGGCUCAUCGAUACGCAUAACAGUGC-- (((((((..(((((.-(((-(.(((((((((((...))))....))---))))).)))).......((((........)))).........)))))))))))).-- ( -35.80, z-score = -1.33, R) >droYak2.chrX 1416772 100 - 21770863 AACUGUUAGGUGCGC-GCAGCGUAGCAGAGCACUCGCUGUUAGGAC---UGUUAAGUGCUCUGUUCACCGCGGGCAAUAGACUUAUCGAUACGCGCAAUAGUGU-- .(((((....(((((-((.(((.(((((((((((.((.((....))---.))..)))))))))))...)))..))....((....)).....))))))))))..-- ( -34.30, z-score = -0.37, R) >droSec1.super_10 1369173 96 - 3036183 AACUGUUAGGUGCGC-GCAGCGUAGCAGAGCACUCGCUGUUAAGUCUGAUGUUAAGUGCUCUGUUCGGCGCGGGCAAUCGACUUAUCGAUACA-AUGA-------- .........((.(((-((..((.(((((((((((.(..((((....))))..).)))))))))))))))))).)).(((((....)))))...-....-------- ( -34.90, z-score = -1.35, R) >droSim1.chrX_random 1032727 96 - 5698898 AACUGUUAGGUGCGC-GCAGCGUAGCAGAGCACUUGCUGUUAAGUCUGAUGUUAAGUGCUCUGUUCGGCGCGGGCAAUCGACUUAUCGAUACA-GUGA-------- .(((((...((.(((-((..((.(((((((((((((..((((....))))..)))))))))))))))))))).)).(((((....))))))))-))..-------- ( -42.80, z-score = -3.62, R) >dp4.chrXL_group1a 4725078 105 + 9151740 -AACAGUGCGUGAACAGUGGGCUGUUAGUGGGCUGCUAAUCGCAGCUGUUGUUAGUUUUGGUGUUACACAGCACUGAUUCCUUCAUCGAUAUUAUCGAUGCAACAA -..(((((((((((((.(..((((.(((..(.((((.....)))))..))).))).)..).)))).))).)))))).......(((((((...)))))))...... ( -35.80, z-score = -2.17, R) >droPer1.super_11 1841150 105 + 2846995 -AACAGUGCGUGAACAGUGGGCUGUUAGCGGGCUGCUAAUCGCAGCUGUUGUUAGUUUUGGUGUUACACAGCACUGAUUCCUUCAUCGAUAUUAUCGAUGCAACAA -..(((((((((((((.(..((((.((((((.((((.....)))))))))).))).)..).)))).))).)))))).......(((((((...)))))))...... ( -38.70, z-score = -2.97, R) >consensus AACUGUUAGGUGCGC_GCAGCGUAGCAGAGCACUCGCUGUUAAGUCUGAUGUUAAGUGCUCUGUUCGCCGCGGGCAAUCGACUUAUCGAUACA_GUGA________ ...((((...(((...))).((((((((((((........................)))))))))....)))))))((((......))))................ (-12.95 = -9.29 + -3.66)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:06:44 2011