| Sequence ID | dm3.chrX |
|---|---|
| Location | 1,466,332 – 1,466,422 |
| Length | 90 |
| Max. P | 0.999692 |

| Location | 1,466,332 – 1,466,422 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 70.90 |
| Shannon entropy | 0.44390 |
| G+C content | 0.50215 |
| Mean single sequence MFE | -26.81 |
| Consensus MFE | -17.40 |
| Energy contribution | -15.78 |
| Covariance contribution | -1.62 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.00 |
| SVM RNA-class probability | 0.978613 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chrX 1466332 90 + 22422827 GCAUGCGUUGCAUGCGUGUGCGUGCAUCCUUUUUUCGGUUUUCGGGGGUUGUUUUGUUCACGAGGACACGCACACACGCACACAAACACA ((((((...))))))(((((((((...........(((((((((.(..(......)..).))))))).))....)))))))))....... ( -32.26, z-score = -2.10, R) >droSec1.super_10 1286841 76 + 3036183 ------GUUGCAUGCGUGUGCGUGCAUCCUUUUUUCGGUUUUCGGGGGUUGUUUUGUUCACGAGGACACGCACACACGCACA-------- ------......((((((((.(((..(((((((..((.....))..)).............)))))..))).))))))))..-------- ( -24.41, z-score = -1.33, R) >droSim1.chrX 1084768 76 + 17042790 ------GUUGCAUGCGUGUGCGUGCAUCCUUUUUUCGGUUUUCGGGGGUUGUUUUGUUCACGAGGACACGCACACACGCACA-------- ------......((((((((.(((..(((((((..((.....))..)).............)))))..))).))))))))..-------- ( -24.41, z-score = -1.33, R) >anoGam1.chrX 8353933 83 + 22145176 GCAUGCACUGUGUGCGUGUGUGUGUGUAUUCUUUUUUGCUUCUCUCCUAUCUUUUGACAACACACACACACAUACACACACUG------- ........((((((.((((((((((((..((........................))..)))))))))))).)))))).....------- ( -26.16, z-score = -3.43, R) >consensus ______GUUGCAUGCGUGUGCGUGCAUCCUUUUUUCGGUUUUCGGGGGUUGUUUUGUUCACGAGGACACGCACACACGCACA________ ........(((..(.(((((((((.(((........))).((((.(((........))).))))..))))))))).)))).......... (-17.40 = -15.78 + -1.62)
| Location | 1,466,332 – 1,466,422 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 70.90 |
| Shannon entropy | 0.44390 |
| G+C content | 0.50215 |
| Mean single sequence MFE | -32.30 |
| Consensus MFE | -25.97 |
| Energy contribution | -24.53 |
| Covariance contribution | -1.44 |
| Combinations/Pair | 1.41 |
| Mean z-score | -4.32 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.20 |
| SVM RNA-class probability | 0.999692 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
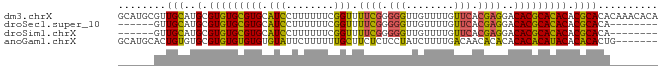
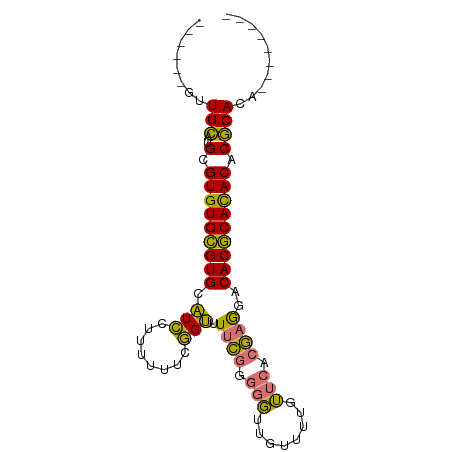
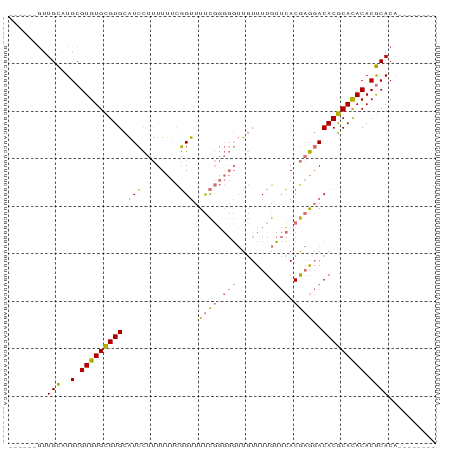
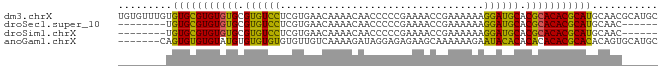
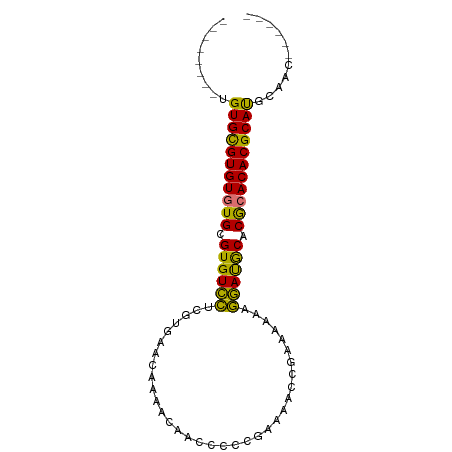
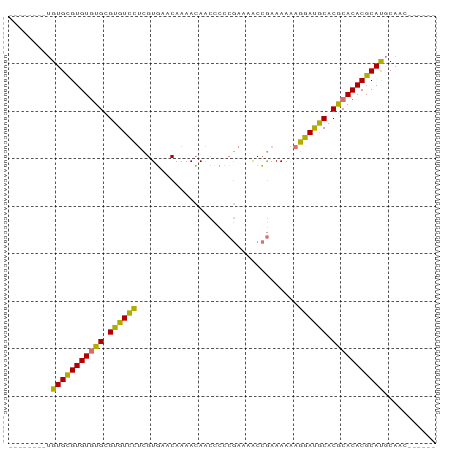
>dm3.chrX 1466332 90 - 22422827 UGUGUUUGUGUGCGUGUGUGCGUGUCCUCGUGAACAAAACAACCCCCGAAAACCGAAAAAAGGAUGCACGCACACGCAUGCAACGCAUGC ((((((.(((((((((((((.(((((((.(....)...........((.....)).....))))))).)))))))))))))))))))... ( -38.10, z-score = -4.25, R) >droSec1.super_10 1286841 76 - 3036183 --------UGUGCGUGUGUGCGUGUCCUCGUGAACAAAACAACCCCCGAAAACCGAAAAAAGGAUGCACGCACACGCAUGCAAC------ --------.(((((((((((.(((((((.(....)...........((.....)).....))))))).))))))))))).....------ ( -28.80, z-score = -4.19, R) >droSim1.chrX 1084768 76 - 17042790 --------UGUGCGUGUGUGCGUGUCCUCGUGAACAAAACAACCCCCGAAAACCGAAAAAAGGAUGCACGCACACGCAUGCAAC------ --------.(((((((((((.(((((((.(....)...........((.....)).....))))))).))))))))))).....------ ( -28.80, z-score = -4.19, R) >anoGam1.chrX 8353933 83 - 22145176 -------CAGUGUGUGUAUGUGUGUGUGUGUUGUCAAAAGAUAGGAGAGAAGCAAAAAAGAAUACACACACACACGCACACAGUGCAUGC -------((.((((((..(((((((((((((((((....))).(........).......))))))))))))))..)))))).))..... ( -33.50, z-score = -4.65, R) >consensus ________UGUGCGUGUGUGCGUGUCCUCGUGAACAAAACAACCCCCGAAAACCGAAAAAAGGAUGCACGCACACGCAUGCAAC______ .........(((((((((((.((((((..................................)))))).)))))))))))........... (-25.97 = -24.53 + -1.44)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:06:23 2011