| Sequence ID | dm3.chrX |
|---|---|
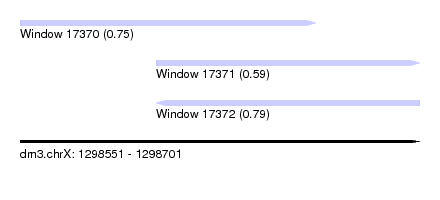
| Location | 1,298,551 – 1,298,701 |
| Length | 150 |
| Max. P | 0.788678 |

| Location | 1,298,551 – 1,298,662 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 94.25 |
| Shannon entropy | 0.09171 |
| G+C content | 0.52678 |
| Mean single sequence MFE | -39.42 |
| Consensus MFE | -36.53 |
| Energy contribution | -36.40 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.93 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.752340 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 1298551 111 + 22422827 CGGCGACUUUUCGCAACUGCAGUUGUAACAAUCGCACCAUUGUAACUGCAAC----ACAGCGAUUGUGAAUGCAAUUGCGACACAAUGGCGGAUGCGGCGCGACUUCUGCGGUUA ..((((....))))((((((((..((.....(((((((.((((....)))).----...((.((((((..(((....))).)))))).)))).)))))....))..)))))))). ( -39.30, z-score = -1.78, R) >droSec1.super_10 1121868 111 + 3036183 CAGCGACUUUUCGCAACUGCAGUUGUAACAAUCGCACCAUUGUAGCUGCAAC----ACAGCGAUUGCGAAUCCGAUUGCGACACAAUGGCGGAUGCGGCGCGACUUCUGCGGUUA ..((((....))))((((((((..((.....((((((((((((.((((....----.))))(.((((((......))))))))))))))....)))))....))..)))))))). ( -39.80, z-score = -1.72, R) >droYak2.chrX 1206780 109 + 21770863 CGGCGACUUUUCGCAACUGCAGUUGUAACAAUCGCACCAUUGUAGCUGCAA------CAGCGAUUGCGAAUCCGAUUGCGACACAAUGGCGGAUGCGGCGUGACUUUUGCGGUUA ..((((....))))((((((((..((.((..((((((((((((.((((...------))))(.((((((......))))))))))))))....))))).)).))..)))))))). ( -38.00, z-score = -1.21, R) >droEre2.scaffold_4644 1282308 115 + 2521924 CGGCGACUUUUCGCAACUGCAGUUGUAACAAUCGCACCAUUGUAGCUGCAACUGCAACAGCGAUUGCGAAUCCGAUUGCGACACAAUGGCGGAUGCGGCGCGACUUCUGCGGUUA ..((((....))))((((((((..((.(((((......))))).((((((.((((....(((((((......)))))))..(.....))))).))))))...))..)))))))). ( -40.60, z-score = -1.12, R) >consensus CGGCGACUUUUCGCAACUGCAGUUGUAACAAUCGCACCAUUGUAGCUGCAAC____ACAGCGAUUGCGAAUCCGAUUGCGACACAAUGGCGGAUGCGGCGCGACUUCUGCGGUUA ..((((....))))((((((((..((.....((((((((((((.((((.........))))(.((((((......))))))))))))))....)))))....))..)))))))). (-36.53 = -36.40 + -0.13)



| Location | 1,298,602 – 1,298,701 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 87.86 |
| Shannon entropy | 0.16306 |
| G+C content | 0.62483 |
| Mean single sequence MFE | -38.63 |
| Consensus MFE | -31.16 |
| Energy contribution | -30.83 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.586457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 1298602 99 + 22422827 CACAGCGAUUGUGAAUGCAAUUGCGACACAAUGGCGGAUGCGGCGCGACUUCUGCGGUUAUCGUCUUGACU-GGGCCAAGACGCGGCCCCCGCAUCCUCC------- ....((.((((((..(((....))).)))))).))((((((((((((.....))))((((......)))).-(((((.......)))))))))))))...------- ( -42.40, z-score = -2.73, R) >droSec1.super_10 1121919 99 + 3036183 CACAGCGAUUGCGAAUCCGAUUGCGACACAAUGGCGGAUGCGGCGCGACUUCUGCGGUUAUCGUCCUGACU-GGGCCAAGACGCCGCCCCCGCAUCCUCC------- ....(((((((......)))))))........((.((((((((.(((.(.(((..((((...((....)).-.)))).))).).)))..)))))))).))------- ( -35.50, z-score = -0.76, R) >droYak2.chrX 1206830 106 + 21770863 -ACAGCGAUUGCGAAUCCGAUUGCGACACAAUGGCGGAUGCGGCGUGACUUUUGCGGUUAUCGUCCUGACUUGGGCCAAGACGCCGCCCCCGCCCCCUGCCUCCUCC -...(((((((......)))))))((..((..(((((..(((((((((((.....))))...((((......))))....)))))))..)))))...))..)).... ( -38.00, z-score = -1.57, R) >consensus CACAGCGAUUGCGAAUCCGAUUGCGACACAAUGGCGGAUGCGGCGCGACUUCUGCGGUUAUCGUCCUGACU_GGGCCAAGACGCCGCCCCCGCAUCCUCC_______ ....(((((((......))))))).........((((..((((((((.....))).(((...((((......))))...))))))))..)))).............. (-31.16 = -30.83 + -0.33)



| Location | 1,298,602 – 1,298,701 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 87.86 |
| Shannon entropy | 0.16306 |
| G+C content | 0.62483 |
| Mean single sequence MFE | -42.90 |
| Consensus MFE | -37.65 |
| Energy contribution | -37.65 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.88 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.788678 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 1298602 99 - 22422827 -------GGAGGAUGCGGGGGCCGCGUCUUGGCCC-AGUCAAGACGAUAACCGCAGAAGUCGCGCCGCAUCCGCCAUUGUGUCGCAAUUGCAUUCACAAUCGCUGUG -------...((((((((((....(((((((((..-.)))))))))....))((.......)).))))))))((.((((((..((....))...)))))).)).... ( -43.30, z-score = -2.62, R) >droSec1.super_10 1121919 99 - 3036183 -------GGAGGAUGCGGGGGCGGCGUCUUGGCCC-AGUCAGGACGAUAACCGCAGAAGUCGCGCCGCAUCCGCCAUUGUGUCGCAAUCGGAUUCGCAAUCGCUGUG -------...((((((((.((((((((((((((..-.)))))))))....)))).(....).).))))))))((.((((((...(.....)...)))))).)).... ( -41.60, z-score = -0.96, R) >droYak2.chrX 1206830 106 - 21770863 GGAGGAGGCAGGGGGCGGGGGCGGCGUCUUGGCCCAAGUCAGGACGAUAACCGCAAAAGUCACGCCGCAUCCGCCAUUGUGUCGCAAUCGGAUUCGCAAUCGCUGU- .......((((.((((((..(((((((((((((....))))))))(((..........)))..)))))..)))))((((((...(.....)...))))))).))))- ( -43.80, z-score = -1.27, R) >consensus _______GGAGGAUGCGGGGGCGGCGUCUUGGCCC_AGUCAGGACGAUAACCGCAGAAGUCGCGCCGCAUCCGCCAUUGUGUCGCAAUCGGAUUCGCAAUCGCUGUG ..........((((((((((....(((((((((....)))))))))....))((.......)).))))))))((.((((((.............)))))).)).... (-37.65 = -37.65 + 0.01)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:05:58 2011