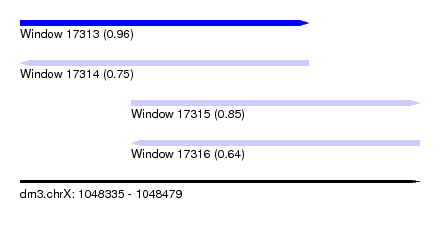
| Sequence ID | dm3.chrX |
|---|---|
| Location | 1,048,335 – 1,048,479 |
| Length | 144 |
| Max. P | 0.964693 |

| Location | 1,048,335 – 1,048,439 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 97.43 |
| Shannon entropy | 0.03532 |
| G+C content | 0.44520 |
| Mean single sequence MFE | -29.77 |
| Consensus MFE | -28.31 |
| Energy contribution | -27.87 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.74 |
| SVM RNA-class probability | 0.964693 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
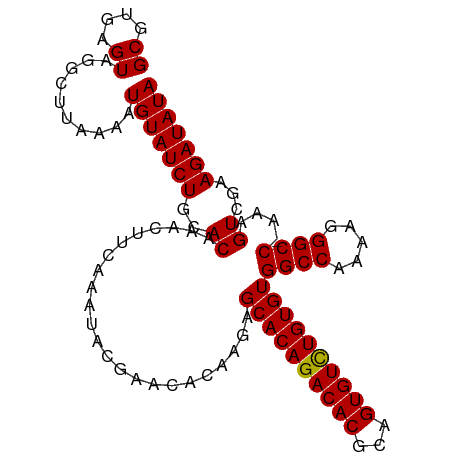
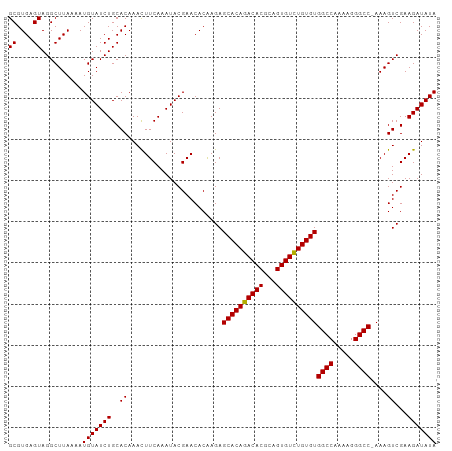
>dm3.chrX 1048335 104 + 22422827 GCGUGAGUAGGCUUAAAAUGUAUCUGCACAAAUUUCAAAUACGAACACAAGAGCACAGACACGCAGUGUUUGUGUGGCCAAAAGGGCCCAAAGUCGAAGAUAUA ((....))..........(((((((..((.......................((((((((((...))))))))))((((.....))))....))...))))))) ( -27.00, z-score = -1.57, R) >droSim1.chrX 838168 103 + 17042790 GCGUGAGUAGGCUUAAAAUGUAUCUGCACAAACUUCAAAUACGAACACGAGAGCACAGACACGCAGUGUCUGUGUGGCCAAAAGGGCC-AAAGUCGAAGAUAUA ((....))..........(((((((........(((......)))..(((..((((((((((...))))))))))((((.....))))-....))).))))))) ( -31.30, z-score = -2.53, R) >droSec1.super_10 878090 103 + 3036183 GCGUGAGUAGGCUUAAAAUGUAUCUGCACAAACUUCAAAUACGAACACAAGAGCACAGACACGCAGUGUCUGUGUGGCCAAAAGGGCC-AAAGUCGAAGAUAUA ..(((((...((.......))..)).)))...((((................((((((((((...))))))))))((((.....))))-......))))..... ( -31.00, z-score = -2.67, R) >consensus GCGUGAGUAGGCUUAAAAUGUAUCUGCACAAACUUCAAAUACGAACACAAGAGCACAGACACGCAGUGUCUGUGUGGCCAAAAGGGCC_AAAGUCGAAGAUAUA ..(((((...((.......))..)).)))...((((................((((((((((...))))))))))((((.....)))).......))))..... (-28.31 = -27.87 + -0.44)

| Location | 1,048,335 – 1,048,439 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 97.43 |
| Shannon entropy | 0.03532 |
| G+C content | 0.44520 |
| Mean single sequence MFE | -26.00 |
| Consensus MFE | -24.64 |
| Energy contribution | -24.53 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.752778 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
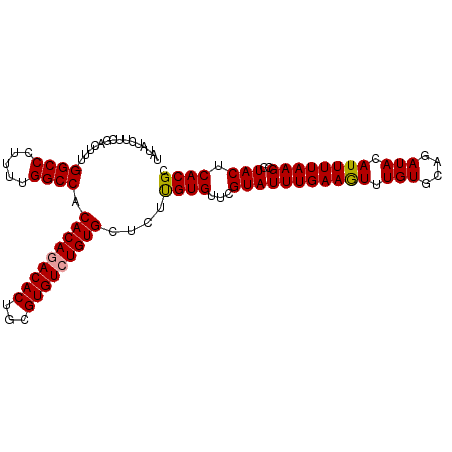
>dm3.chrX 1048335 104 - 22422827 UAUAUCUUCGACUUUGGGCCCUUUUGGCCACACAAACACUGCGUGUCUGUGCUCUUGUGUUCGUAUUUGAAAUUUGUGCAGAUACAUUUUAAGCCUACUCACGC ..............((((((.....)))).))........(((((..((.(((..(((((..((((.........))))..))))).....))).))..))))) ( -23.70, z-score = -1.04, R) >droSim1.chrX 838168 103 - 17042790 UAUAUCUUCGACUUU-GGCCCUUUUGGCCACACAGACACUGCGUGUCUGUGCUCUCGUGUUCGUAUUUGAAGUUUGUGCAGAUACAUUUUAAGCCUACUCACGC ..............(-((((.....)))))(((((((((...)))))))))....((((...(((((((((((.(((....))).))))))))..))).)))). ( -27.50, z-score = -1.79, R) >droSec1.super_10 878090 103 - 3036183 UAUAUCUUCGACUUU-GGCCCUUUUGGCCACACAGACACUGCGUGUCUGUGCUCUUGUGUUCGUAUUUGAAGUUUGUGCAGAUACAUUUUAAGCCUACUCACGC ..............(-((((.....)))))(((((((((...)))))))))....(((((..((((.........))))..))))).................. ( -26.80, z-score = -1.60, R) >consensus UAUAUCUUCGACUUU_GGCCCUUUUGGCCACACAGACACUGCGUGUCUGUGCUCUUGUGUUCGUAUUUGAAGUUUGUGCAGAUACAUUUUAAGCCUACUCACGC ................((((.....)))).(((((((((...)))))))))....((((...(((((((((((.(((....))).))))))))..))).)))). (-24.64 = -24.53 + -0.11)

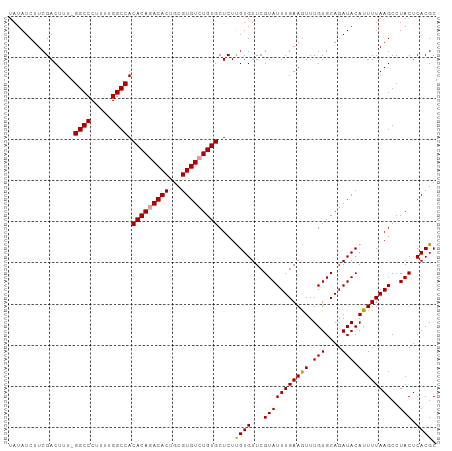
| Location | 1,048,375 – 1,048,479 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 97.43 |
| Shannon entropy | 0.03532 |
| G+C content | 0.52262 |
| Mean single sequence MFE | -32.61 |
| Consensus MFE | -31.34 |
| Energy contribution | -31.12 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.96 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.850746 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 1048375 104 + 22422827 ACGAACACAAGAGCACAGACACGCAGUGUUUGUGUGGCCAAAAGGGCCCAAAGUCGAAGAUAUAGCCGCACUUUGCAAAUCGUGACUGGGCAAAGGCACUUACC .(((........((((((((((...))))))))))((((.....)))).....)))...........((.((((((.............))))))))....... ( -29.92, z-score = -0.92, R) >droSim1.chrX 838208 103 + 17042790 ACGAACACGAGAGCACAGACACGCAGUGUCUGUGUGGCCAAAAGGGCC-AAAGUCGAAGAUAUAGCCGCACUUUGCAAAUCGUGACUGGGCAAAGGCACCUACC .....(((((..((((((((((...))))))))))((((.....))))-(((((((..........)).))))).....)))))...((((....)).)).... ( -34.30, z-score = -1.98, R) >droSec1.super_10 878130 103 + 3036183 ACGAACACAAGAGCACAGACACGCAGUGUCUGUGUGGCCAAAAGGGCC-AAAGUCGAAGAUAUAGCCGCACUUUGCAAAUCGUGACUGGGCAAAGGCACCUACC .(((........((((((((((...))))))))))((((.....))))-....)))......(((..((.((((((.............))))))))..))).. ( -33.62, z-score = -1.95, R) >consensus ACGAACACAAGAGCACAGACACGCAGUGUCUGUGUGGCCAAAAGGGCC_AAAGUCGAAGAUAUAGCCGCACUUUGCAAAUCGUGACUGGGCAAAGGCACCUACC .(((........((((((((((...))))))))))((((.....)))).....)))...........((.((((((.............))))))))....... (-31.34 = -31.12 + -0.22)

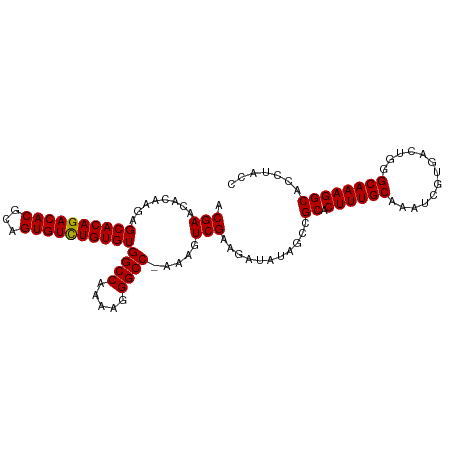

| Location | 1,048,375 – 1,048,479 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 97.43 |
| Shannon entropy | 0.03532 |
| G+C content | 0.52262 |
| Mean single sequence MFE | -33.30 |
| Consensus MFE | -31.53 |
| Energy contribution | -32.20 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.644235 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 1048375 104 - 22422827 GGUAAGUGCCUUUGCCCAGUCACGAUUUGCAAAGUGCGGCUAUAUCUUCGACUUUGGGCCCUUUUGGCCACACAAACACUGCGUGUCUGUGCUCUUGUGUUCGU (((....)))...((.(((.((((...((((((((.(((........)))))))))((((.....)))).......))...)))).))).))............ ( -28.10, z-score = 0.04, R) >droSim1.chrX 838208 103 - 17042790 GGUAGGUGCCUUUGCCCAGUCACGAUUUGCAAAGUGCGGCUAUAUCUUCGACUUU-GGCCCUUUUGGCCACACAGACACUGCGUGUCUGUGCUCUCGUGUUCGU .((((.(((((((((..(((....))).)))))).))).)))).....(((...(-((((.....)))))(((((((((...))))))))).........))). ( -35.90, z-score = -1.98, R) >droSec1.super_10 878130 103 - 3036183 GGUAGGUGCCUUUGCCCAGUCACGAUUUGCAAAGUGCGGCUAUAUCUUCGACUUU-GGCCCUUUUGGCCACACAGACACUGCGUGUCUGUGCUCUUGUGUUCGU .((((.(((((((((..(((....))).)))))).))).)))).....(((...(-((((.....)))))(((((((((...))))))))).........))). ( -35.90, z-score = -2.02, R) >consensus GGUAGGUGCCUUUGCCCAGUCACGAUUUGCAAAGUGCGGCUAUAUCUUCGACUUU_GGCCCUUUUGGCCACACAGACACUGCGUGUCUGUGCUCUUGUGUUCGU .((((.(((((((((..(((....))).)))))).))).)))).....(((.....((((.....)))).(((((((((...))))))))).........))). (-31.53 = -32.20 + 0.67)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:05:12 2011