| Sequence ID | dm3.chrX |
|---|---|
| Location | 846,764 – 846,877 |
| Length | 113 |
| Max. P | 0.902088 |

| Location | 846,764 – 846,877 |
|---|---|
| Length | 113 |
| Sequences | 8 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 92.38 |
| Shannon entropy | 0.15499 |
| G+C content | 0.45814 |
| Mean single sequence MFE | -22.47 |
| Consensus MFE | -20.30 |
| Energy contribution | -20.30 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.902088 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
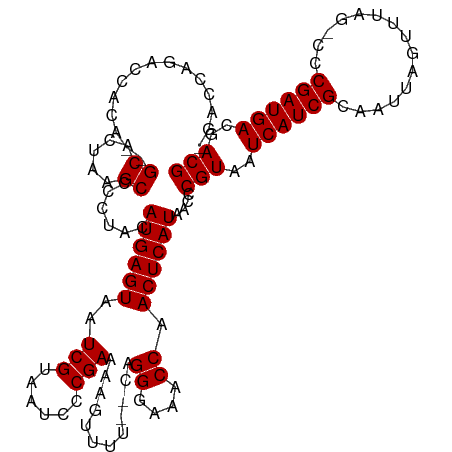
>dm3.chrX 846764 113 - 22422827 GAACCAGACCACA--GCACUAAGCCCCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUU--CAGGGAAACCAACUCAUAACCCGUAAUCAUCGCAAUUAGUUUAG-CCCGAUGACGACG .............--((.....))......((((((..(((......)))........--..((....)).))))))....(((..((((((............-..))))))..))) ( -20.24, z-score = -1.80, R) >droSim1.chrX 681298 113 - 17042790 GAACCAGACCACA--GCACUAAGCCCCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUU--CAGGGAAACCAACUCAUAACCCGUAAUCAUCGCAAUUAGUUUAG-CCCGAUGACGACG .............--((.....))......((((((..(((......)))........--..((....)).))))))....(((..((((((............-..))))))..))) ( -20.24, z-score = -1.80, R) >droSec1.super_10 683453 113 - 3036183 GAACCAGACCACA--GCACUAAGCCCCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUU--CAGGGAAACCAACUCAUAACCCGUAAUCAUCGCAAUUAGUUUAG-CCCGAUGACGACG .............--((.....))......((((((..(((......)))........--..((....)).))))))....(((..((((((............-..))))))..))) ( -20.24, z-score = -1.80, R) >droYak2.chrX 763214 113 - 21770863 AGACCAGACCGCA--GCUCUAAGCCCCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUU--CAGGGAAACCAACUCAUAACCCGUAAUCAUCGCAAUUAGUUUAG-CUCGAUGACGACG .............--((.....))......((((((..(((......)))........--..((....)).))))))....(((..((((((.....((.....-))))))))..))) ( -20.80, z-score = -0.87, R) >droEre2.scaffold_4644 817093 113 - 2521924 AGGCCAGACCACA--GCUCUAAGCCCCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUU--CAGGGAAACCAACUCAUAACCCGUAAUCAUCGCAAUUAGUUUAG-CACGAUGACGACG .(((.(((.....--..)))..))).....((((((..(((......)))........--..((....)).))))))....(((..((((((............-..))))))..))) ( -23.74, z-score = -1.90, R) >droAna3.scaffold_12929 607882 113 - 3277472 -----AGACCACACAGCACUAAGCUCCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUUGCUUGGGAAACCAACUCAUAACCCGUAAUCAUCGUAAUUAGUUUAGCCCCGAUGACGACG -----.........(((.....))).....((((((....((((..(......)..)))).(((....)))))))))....(((..((((((...............))))))..))) ( -22.86, z-score = -1.74, R) >dp4.chrXL_group1a 3804632 115 + 9151740 -GAACAGACCACAGCGCACUAAGCUUCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUU-GCAGGGAAACCAACUCAUAACCCGUAAUCAUCGCAAUUAGUUUGC-CCCGAUGACGACG -....(((....(((.......))))))..((((((....((((..(......)..))-)).((....)).))))))....(((..(((((((((.....))))-...)))))..))) ( -25.80, z-score = -2.28, R) >droPer1.super_11 867011 115 + 2846995 -GAACAGACCACAGCGCACUAAGCUUCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUU-GCAGGGAAACCAACUCAUAACCCGUAAUCAUCGCAAUUAGUUUGC-CCCGAUGACGACG -....(((....(((.......))))))..((((((....((((..(......)..))-)).((....)).))))))....(((..(((((((((.....))))-...)))))..))) ( -25.80, z-score = -2.28, R) >consensus _GACCAGACCACA__GCACUAAGCCCCUACAUGAGUAAUCGUAAUCCCGAAAAGUUUU__CAGGGAAACCAACUCAUAACCCGUAAUCAUCGCAAUUAGUUUAG_CCCGAUGACGACG ...............((.....))......((((((..(((......)))............((....)).))))))....(((..((((((...............))))))..))) (-20.30 = -20.30 + -0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:04:19 2011