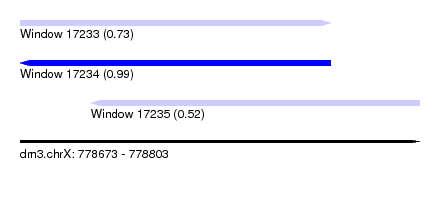

| Sequence ID | dm3.chrX |
|---|---|
| Location | 778,673 – 778,803 |
| Length | 130 |
| Max. P | 0.994719 |

| Location | 778,673 – 778,774 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 75.25 |
| Shannon entropy | 0.38642 |
| G+C content | 0.51918 |
| Mean single sequence MFE | -30.70 |
| Consensus MFE | -24.14 |
| Energy contribution | -24.32 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.08 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.733830 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 778673 101 + 22422827 CUUUCGCCAAUUUCCCGAGUGGGAAAUUCGGUUCGAAGGACCAAGGUCCUUCAGGUGGGACUUUUUCGAACGACGAACGUG--UUUUUUGGCGAGCAUUCGAG ..((((((((((((((....))))))....(((((((((((....)))))))..((.(..(......)..).))))))...--....))))))))........ ( -36.30, z-score = -1.80, R) >droSim1.chrX 625208 103 + 17042790 CUUUCGCCAAUUUCCCGAGUGGGAAAAUCGGUUCGAAGGACCAAGGUCCUUCUGGUGGGACUUUUUCAAACGAUGAGCGUGCGUUCUUGGGCAAACAUUCGAG .(..(((((.((((((....))))))........(((((((....))))))))))))..)..........(((......(((.(....).))).....))).. ( -33.10, z-score = -0.90, R) >droYak2.chrX 697492 80 + 21770863 CUUUCGGCAAUUUCCCGAGUGGGAAAAUCGCUUCGCAGGACCAAGGUCCUUCAGGUGGGACUUUUUCGAACGACGACGAG----------------------- ..(((((.((..((((((((((.....)))))).(.(((((....))))).)....))))..)).)))))..........----------------------- ( -23.50, z-score = 0.01, R) >droEre2.scaffold_4644 741442 85 + 2521924 CUUUCAGCAAUUUCCCGAGUGGGAAAAUCGCUGCGAAGGACCAAGGUCCUUCAGGUGGGACUUUUUCGAACGGCGAGCAAUCGAG------------------ ......((..((((((....)))))).((((((.(((((((....)))))))...((..(...)..))..)))))))).......------------------ ( -29.90, z-score = -1.64, R) >consensus CUUUCGCCAAUUUCCCGAGUGGGAAAAUCGCUUCGAAGGACCAAGGUCCUUCAGGUGGGACUUUUUCGAACGACGAGCAUG__UU__________________ .(..((((..((((((....))))))........(((((((....))))))).))))..).....(((.....)))........................... (-24.14 = -24.32 + 0.19)

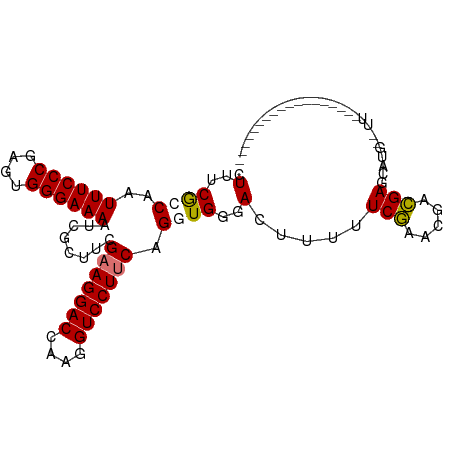
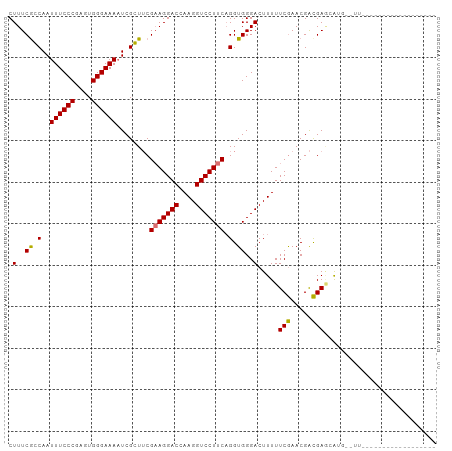
| Location | 778,673 – 778,774 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 75.25 |
| Shannon entropy | 0.38642 |
| G+C content | 0.51918 |
| Mean single sequence MFE | -29.67 |
| Consensus MFE | -25.17 |
| Energy contribution | -24.55 |
| Covariance contribution | -0.62 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.73 |
| SVM RNA-class probability | 0.994719 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 778673 101 - 22422827 CUCGAAUGCUCGCCAAAAAA--CACGUUCGUCGUUCGAAAAAGUCCCACCUGAAGGACCUUGGUCCUUCGAACCGAAUUUCCCACUCGGGAAAUUGGCGAAAG .........(((((((....--...(((((..((((((.....))......(((((((....))))))))))))))))(((((....))))).)))))))... ( -34.80, z-score = -3.51, R) >droSim1.chrX 625208 103 - 17042790 CUCGAAUGUUUGCCCAAGAACGCACGCUCAUCGUUUGAAAAAGUCCCACCAGAAGGACCUUGGUCCUUCGAACCGAUUUUCCCACUCGGGAAAUUGGCGAAAG ..(((..((.(((........))).))...)))((((..............(((((((....)))))))...((((((((((.....)))))))))))))).. ( -29.30, z-score = -1.45, R) >droYak2.chrX 697492 80 - 21770863 -----------------------CUCGUCGUCGUUCGAAAAAGUCCCACCUGAAGGACCUUGGUCCUGCGAAGCGAUUUUCCCACUCGGGAAAUUGCCGAAAG -----------------------....(((((((..((..((((((........))).)))..))..)))).((((((((((.....)))))))))))))... ( -23.80, z-score = -1.23, R) >droEre2.scaffold_4644 741442 85 - 2521924 ------------------CUCGAUUGCUCGCCGUUCGAAAAAGUCCCACCUGAAGGACCUUGGUCCUUCGCAGCGAUUUUCCCACUCGGGAAAUUGCUGAAAG ------------------...((((..(((.....)))...))))......(((((((....))))))).((((((((((((.....)))))))))))).... ( -30.80, z-score = -3.32, R) >consensus __________________AA__CACGCUCGUCGUUCGAAAAAGUCCCACCUGAAGGACCUUGGUCCUUCGAACCGAUUUUCCCACUCGGGAAAUUGCCGAAAG .................................((((..............(((((((....)))))))...((((.((((((....)))))))))))))).. (-25.17 = -24.55 + -0.62)

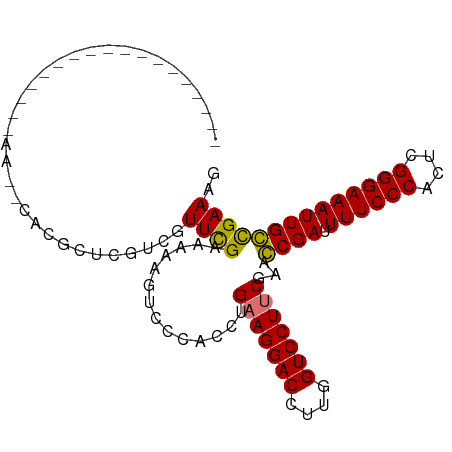
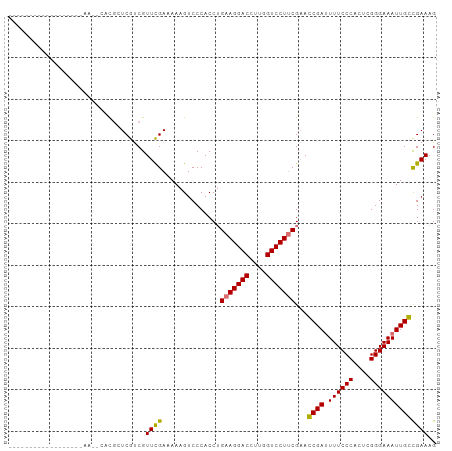
| Location | 778,696 – 778,803 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 70.06 |
| Shannon entropy | 0.48026 |
| G+C content | 0.48823 |
| Mean single sequence MFE | -22.23 |
| Consensus MFE | -13.57 |
| Energy contribution | -13.33 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.17 |
| Mean z-score | -0.95 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.519487 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 778696 107 - 22422827 UGUACAGUCCCAAUGCUAAUAUUCCUUCGCUCGAAUGCUCGCCAAAAAA--CACGUUCGUCGUUCGAAAAAGUCCCACCUGAAGGACCUUGGUCCUUCGAACCGAAUUU .........................((((.(((((((..((........--......)).)))))))............((((((((....))))))))...))))... ( -20.44, z-score = -0.71, R) >droSim1.chrX 625231 109 - 17042790 UGUACAGCCCCAAUGAUACUGUUCCUUCGCUCGAAUGUUUGCCCAAGAACGCACGCUCAUCGUUUGAAAAAGUCCCACCAGAAGGACCUUGGUCCUUCGAACCGAUUUU .(((((.......)).))).((((.((((..(((..((.(((........))).))...)))..))))............(((((((....)))))))))))....... ( -24.10, z-score = -1.12, R) >droYak2.chrX 697515 87 - 21770863 GAUAUAUCCAC----UAGAUC-CACU--GCUCUCUUGCUCG---------------UCGUCGUUCGAAAAAGUCCCACCUGAAGGACCUUGGUCCUGCGAAGCGAUUUU ...........----......-....--((......))(((---------------(..((((..((..((((((........))).)))..))..)))).)))).... ( -15.90, z-score = 0.48, R) >droEre2.scaffold_4644 741465 97 - 2521924 CGCACAGCCACGAUUUAAUUCGUACU--GUUCUCUUGCUCG----------AUUGCUCGCCGUUCGAAAAAGUCCCACCUGAAGGACCUUGGUCCUUCGCAGCGAUUUU (((((((..((((......)))).))--)).....(((..(----------(((..(((.....)))...))))......(((((((....)))))))))))))..... ( -28.50, z-score = -2.45, R) >consensus UGUACAGCCACAAUGUAAAUCUUACU__GCUCGAAUGCUCG__________CACGCUCGUCGUUCGAAAAAGUCCCACCUGAAGGACCUUGGUCCUUCGAACCGAUUUU ............................((......))..................(((.....))).((((((......(((((((....))))))).....)))))) (-13.57 = -13.33 + -0.25)

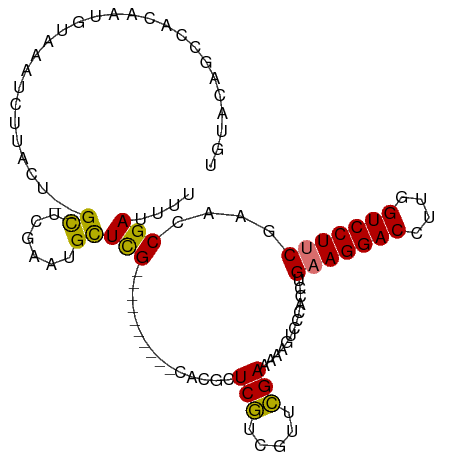
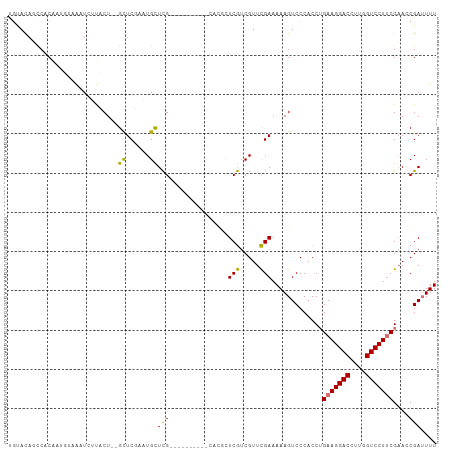
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:04:06 2011