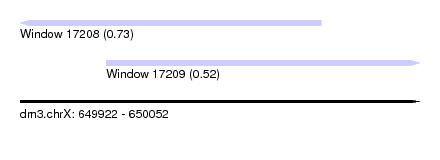

| Sequence ID | dm3.chrX |
|---|---|
| Location | 649,922 – 650,052 |
| Length | 130 |
| Max. P | 0.727437 |

| Location | 649,922 – 650,020 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 88.19 |
| Shannon entropy | 0.20866 |
| G+C content | 0.53760 |
| Mean single sequence MFE | -38.24 |
| Consensus MFE | -28.64 |
| Energy contribution | -30.20 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.727437 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
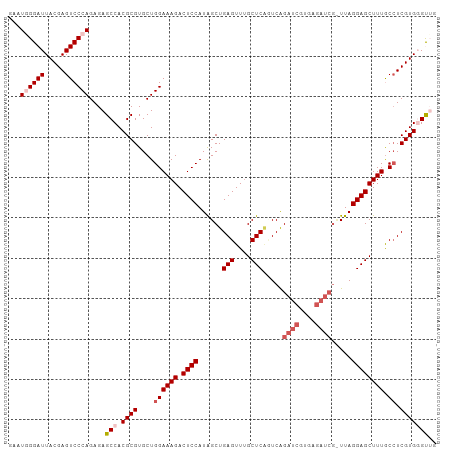
>dm3.chrX 649922 98 - 22422827 UUAUGGGAUUACGAGUCCCAGAGAGCCACGCGUGCUGGAAAGACUCCAUAGCUGAGUUUGCUCCGUCAGAUCGC--------UUAGGAGCUUUGCCUCGUGGGUUG ...((((((.....))))))...((((.((((....((((((.((((..(((.((.((((......))))))))--------)..)))))))).)).)))))))). ( -32.90, z-score = -0.86, R) >droSim1.chrX 536502 105 - 17042790 GAAUGGGAUUACGAGUCCCAGAGAGCCACGCGUGCUGGAAAGACUCCAUAGCUGAGUUUGCUCGUUCAGAUCAUGAGAUCGCUUAGGAGCUUUACGUCGUGCGUG- ...((((((.....))))))......((((((((.((.((((.((((..(((.(((....))).....((((....)))))))..)))))))).)).))))))))- ( -37.20, z-score = -1.86, R) >droSec1.super_10 499441 105 - 3036183 GAAUGGGAUUACGAGUCCCAGAGAGCCACGCGUGCUGGAAAGACUCCAUAGCUGAGUUUGCUCGUUCAGAUCGUGAGAUCGUUUAGGAGCUUUGCCUCGUGCGUG- ...((((((.....))))))......((((((((..((((((.((((..(((.(((....))).....((((....)))))))..)))))))).)).))))))))- ( -39.70, z-score = -2.49, R) >droYak2.chrX 575926 106 - 21770863 GAAUAGGAUUACGAGUCCCAGAGAGCCACGCGUGCUGGAAAGACUCCAUGGCUGAGUUUGCUCACUCAGAUCGCGAGAUCGUUUAGGAGCUUUGCCUCGUGGGCUG .....((((.....)))).....((((.((((....((((((.((((.((..((((....))))..))((((....)))).....)))))))).)).)))))))). ( -39.60, z-score = -1.74, R) >droEre2.scaffold_4644 614221 105 - 2521924 CAAUAGGAUUACGAGUCCCAGAGAGCCACGCGUGCUGGAAAGACUCCAUAGCUGAGUUCGCUCAGUCGGAUCGUGAGAUCG-UUAGGAGCUUUACCUCGUGGGCUG .....((((.....)))).....((((.((((....((((((.((((...((((((....))))))..((((....)))).-...)))))))).)).)))))))). ( -41.80, z-score = -2.82, R) >consensus GAAUGGGAUUACGAGUCCCAGAGAGCCACGCGUGCUGGAAAGACUCCAUAGCUGAGUUUGCUCAGUCAGAUCGUGAGAUCG_UUAGGAGCUUUGCCUCGUGGGUUG ...((((((.....))))))....(((.((((....((((((.((((......(((....))).....((((....)))).....)))))))).)).))))))).. (-28.64 = -30.20 + 1.56)


| Location | 649,950 – 650,052 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 90.27 |
| Shannon entropy | 0.17757 |
| G+C content | 0.52378 |
| Mean single sequence MFE | -31.26 |
| Consensus MFE | -28.54 |
| Energy contribution | -28.30 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.04 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.520787 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
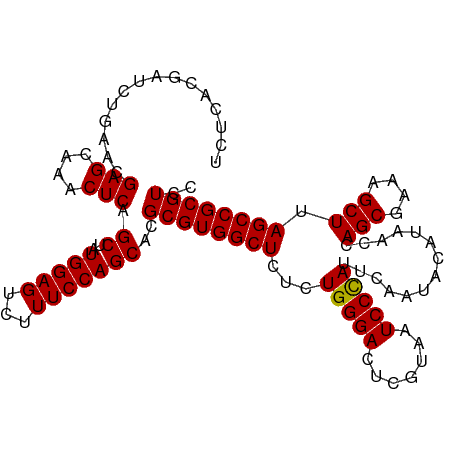
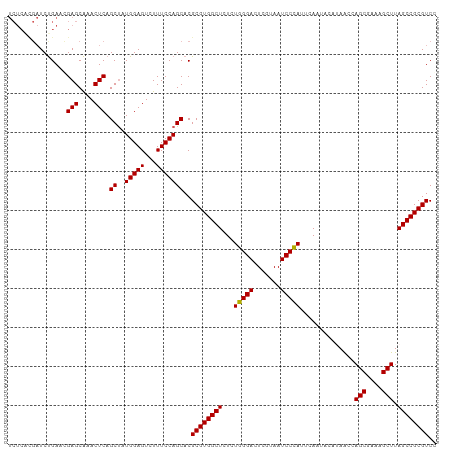
>dm3.chrX 649950 102 + 22422827 --------UCUGACGGAGCAAACUCAGCUAUGGAGUCUUUCCAGCACGCGUGGCUCUCUGGGACUCGUAAUCCCAUAAAAUACAUAACCAGCGAAAGCUUAGCCGCGUCU --------...(((((((...((((.......))))..)))).......((((((...(((((.......)))))..............(((....))).))))))))). ( -30.40, z-score = -1.59, R) >droSim1.chrX 536529 110 + 17042790 UCUCAUGAUCUGAACGAGCAAACUCAGCUAUGGAGUCUUUCCAGCACGCGUGGCUCUCUGGGACUCGUAAUCCCAUUCAAUACAUAACCAGCGAAAGCUUAGCCGCGUCC ...............(((....))).(((..((((...)))))))..((((((((...(((((.......)))))..............(((....))).)))))))).. ( -29.40, z-score = -1.11, R) >droSec1.super_10 499468 110 + 3036183 UCUCACGAUCUGAACGAGCAAACUCAGCUAUGGAGUCUUUCCAGCACGCGUGGCUCUCUGGGACUCGUAAUCCCAUUCAAUACAUAACCAGCGAAAGCUUAGCCGCGUCC ...............(((....))).(((..((((...)))))))..((((((((...(((((.......)))))..............(((....))).)))))))).. ( -29.40, z-score = -1.14, R) >droYak2.chrX 575954 110 + 21770863 UCUCGCGAUCUGAGUGAGCAAACUCAGCCAUGGAGUCUUUCCAGCACGCGUGGCUCUCUGGGACUCGUAAUCCUAUUCAAUACAAAGCCAGCGAAAGCUUAGCCGCGUCG ....((...((((((......))))))...(((((...))))))).(((((((((...(((((.......)))))..............(((....))).))))))).)) ( -33.90, z-score = -1.03, R) >droEre2.scaffold_4644 614248 110 + 2521924 UCUCACGAUCCGACUGAGCGAACUCAGCUAUGGAGUCUUUCCAGCACGCGUGGCUCUCUGGGACUCGUAAUCCUAUUGAAGACAAAGCCAGCGAAAGCUUAGCCGCGUCG .....((((.((.((((((.......(((..((((...))))))).(((.(((((...(((((.......)))))(((....)))))))))))...)))))).)).)))) ( -33.20, z-score = -0.33, R) >consensus UCUCACGAUCUGAACGAGCAAACUCAGCUAUGGAGUCUUUCCAGCACGCGUGGCUCUCUGGGACUCGUAAUCCCAUUCAAUACAUAACCAGCGAAAGCUUAGCCGCGUCC ...............(((....))).((..(((((...)))))))..((((((((...(((((.......)))))..............(((....))).)))))))).. (-28.54 = -28.30 + -0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:03:44 2011