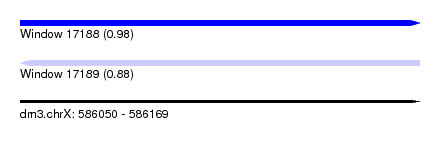

| Sequence ID | dm3.chrX |
|---|---|
| Location | 586,050 – 586,169 |
| Length | 119 |
| Max. P | 0.980291 |

| Location | 586,050 – 586,169 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.65 |
| Shannon entropy | 0.12489 |
| G+C content | 0.35013 |
| Mean single sequence MFE | -33.50 |
| Consensus MFE | -25.00 |
| Energy contribution | -25.04 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.25 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.04 |
| SVM RNA-class probability | 0.980291 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
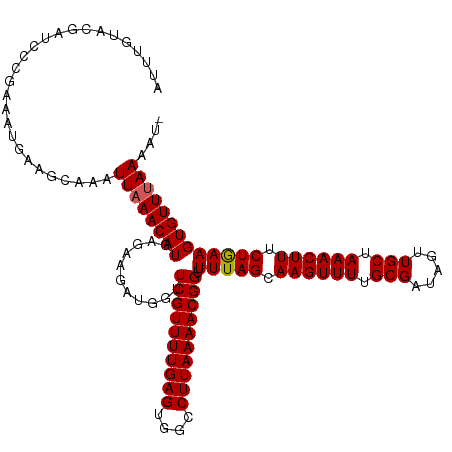
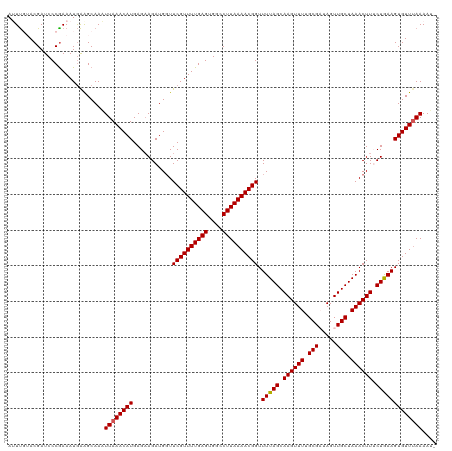
>dm3.chrX 586050 119 + 22422827 AUUCGGACGAUCCCGAAAUAGGGCAAAUUAAACAUGAGAAGAUGGUUCGUUUUGAGUGGCCUUAAAACGGUUUUAGCAAGUUUUGCGAUAGUUGCUAAACUUUCUGAAGUGUUUAAAAU- .(((((......)))))..........((((((((....(((.(((((((((((((....))))))))))..(((((((.(........).)))))))))).)))...))))))))...- ( -32.20, z-score = -2.33, R) >droSim1.chrX 471277 119 + 17042790 AUUUGGACGAUCCCAAAAUGAAGCAAAUUAAACAUGAGAAGAUGGUUCGUUUUGAGUGGCCUUAAAACGGUUUUAGCAAGUUUUGCGAUAGUUGCUAAACUUGCUGAAGUGUUUAAAAU- .(((((......)))))..........((((((((...........((((((((((....)))))))))).((((((((((((.((((...)))).))))))))))))))))))))...- ( -36.10, z-score = -4.05, R) >droSec1.super_10 433331 119 + 3036183 AUUUGUACGAUCCCGAAAUGAAGCAAAUUAAACAUGAGAAGAUGGUUCGUUUUGAGUGGCCUUAAAACGGUUUUAGCAAGUUUUGCGAUAGUUGCUAAACUUGCUGAAGUGUUUAAAAU- .(((((................)))))((((((((...........((((((((((....)))))))))).((((((((((((.((((...)))).))))))))))))))))))))...- ( -32.49, z-score = -3.07, R) >droYak2.chrX 511120 120 + 21770863 AGUUUUACGAUCCCGAAAUGAAGCAAAUUAAACAUGAGUAGAUGCUUCGUUUUGAGUGGCCUUAAAACGGUUUUAGCAAGUUUAGCGUUAGUUGCUAAACUUUCUAAAGUGUUUAAAAAU .((((((((....))...))))))...((((((((...((((((((((((((((((....))))))))))....))))(((((((((.....))))))))).))))..)))))))).... ( -35.00, z-score = -4.01, R) >droEre2.scaffold_4644 549462 120 + 2521924 AGUUUUACGAUCCCGAAAUGAAGCAAUUUAAACAUGAGAAGAUGGUUCGUUUUGAGUGGCCUUAAAACGGUUUUAGCAAGUUUAGCGUUAGUUGCUAAACUUUCUGAAGUGUUAAAAAAU .((((((.(..(((((((((((.((.(((........)))..)).))))))))).).)..).))))))..((((((.((((((((((.....)))))))))).))))))........... ( -31.70, z-score = -2.79, R) >consensus AUUUGUACGAUCCCGAAAUGAAGCAAAUUAAACAUGAGAAGAUGGUUCGUUUUGAGUGGCCUUAAAACGGUUUUAGCAAGUUUUGCGAUAGUUGCUAAACUUUCUGAAGUGUUUAAAAU_ ...........................((((((((...........((((((((((....)))))))))).(((((.((((((.(((.....))).)))))).))))))))))))).... (-25.00 = -25.04 + 0.04)

| Location | 586,050 – 586,169 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.65 |
| Shannon entropy | 0.12489 |
| G+C content | 0.35013 |
| Mean single sequence MFE | -23.96 |
| Consensus MFE | -15.98 |
| Energy contribution | -16.18 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.95 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.882643 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
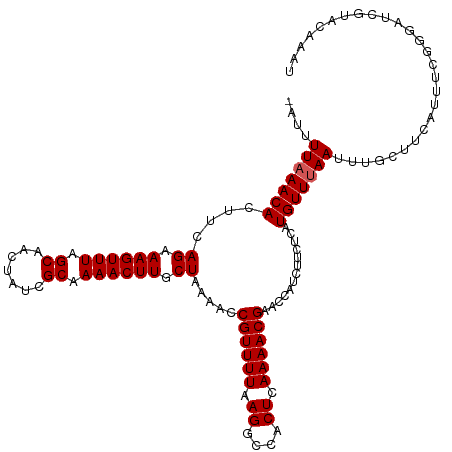
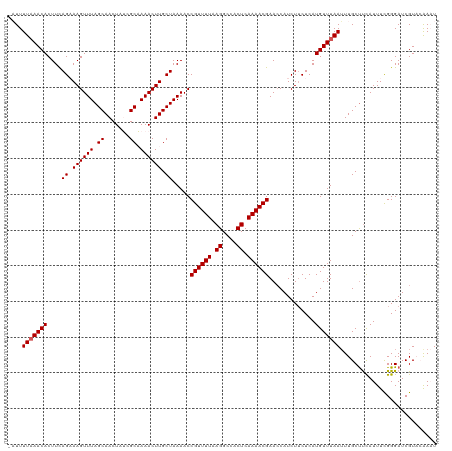
>dm3.chrX 586050 119 - 22422827 -AUUUUAAACACUUCAGAAAGUUUAGCAACUAUCGCAAAACUUGCUAAAACCGUUUUAAGGCCACUCAAAACGAACCAUCUUCUCAUGUUUAAUUUGCCCUAUUUCGGGAUCGUCCGAAU -...(((((((....((((..((((((((............))))))))..((((((.((....)).)))))).......))))..)))))))..........((((((....)))))). ( -22.40, z-score = -2.13, R) >droSim1.chrX 471277 119 - 17042790 -AUUUUAAACACUUCAGCAAGUUUAGCAACUAUCGCAAAACUUGCUAAAACCGUUUUAAGGCCACUCAAAACGAACCAUCUUCUCAUGUUUAAUUUGCUUCAUUUUGGGAUCGUCCAAAU -...(((((((....(((((((((.((.......)).))))))))).....((((((.((....)).)))))).............)))))))..........((((((....)))))). ( -26.90, z-score = -4.20, R) >droSec1.super_10 433331 119 - 3036183 -AUUUUAAACACUUCAGCAAGUUUAGCAACUAUCGCAAAACUUGCUAAAACCGUUUUAAGGCCACUCAAAACGAACCAUCUUCUCAUGUUUAAUUUGCUUCAUUUCGGGAUCGUACAAAU -...(((((((....(((((((((.((.......)).))))))))).....((((((.((....)).)))))).............)))))))........................... ( -23.20, z-score = -3.16, R) >droYak2.chrX 511120 120 - 21770863 AUUUUUAAACACUUUAGAAAGUUUAGCAACUAACGCUAAACUUGCUAAAACCGUUUUAAGGCCACUCAAAACGAAGCAUCUACUCAUGUUUAAUUUGCUUCAUUUCGGGAUCGUAAAACU ........((..(((((.(((((((((.......))))))))).))))).(((((((.((....)).)))))((((((...((....))......)))))).....))....))...... ( -28.20, z-score = -3.97, R) >droEre2.scaffold_4644 549462 120 - 2521924 AUUUUUUAACACUUCAGAAAGUUUAGCAACUAACGCUAAACUUGCUAAAACCGUUUUAAGGCCACUCAAAACGAACCAUCUUCUCAUGUUUAAAUUGCUUCAUUUCGGGAUCGUAAAACU ...............((.(((((((((.......))))))))).)).....((((((.((....)).)))))).((.(((((...(((............)))...))))).))...... ( -19.10, z-score = -1.32, R) >consensus _AUUUUAAACACUUCAGAAAGUUUAGCAACUAUCGCAAAACUUGCUAAAACCGUUUUAAGGCCACUCAAAACGAACCAUCUUCUCAUGUUUAAUUUGCUUCAUUUCGGGAUCGUACAAAU ....(((((((....((.((((((.((.......)).)))))).)).....((((((.((....)).)))))).............)))))))........................... (-15.98 = -16.18 + 0.20)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:03:27 2011