
| Sequence ID | dm3.chrX |
|---|---|
| Location | 519,222 – 519,289 |
| Length | 67 |
| Max. P | 0.957692 |

| Location | 519,222 – 519,289 |
|---|---|
| Length | 67 |
| Sequences | 5 |
| Columns | 69 |
| Reading direction | forward |
| Mean pairwise identity | 82.15 |
| Shannon entropy | 0.30933 |
| G+C content | 0.49846 |
| Mean single sequence MFE | -20.29 |
| Consensus MFE | -16.40 |
| Energy contribution | -16.96 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.941560 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
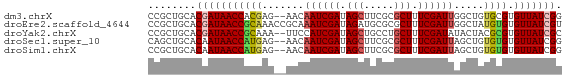
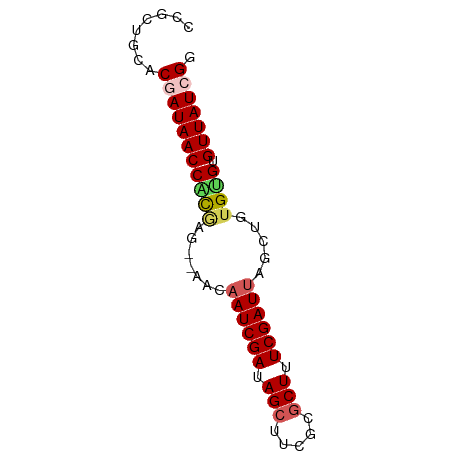
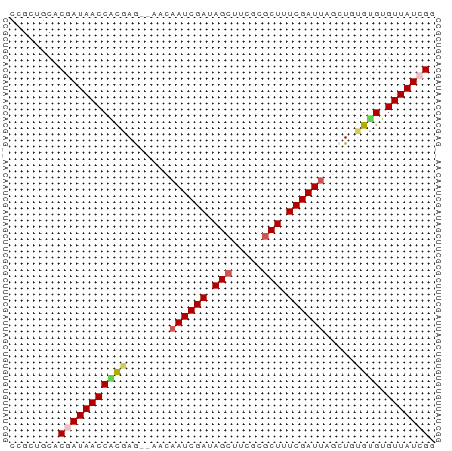
>dm3.chrX 519222 67 + 22422827 CCGCUGCACGAUAACCACGAG--AACAAUCGAUAGCUUCGCGCUUUCGAUUGGCUGUGCGUGUUAUCGG ........(((((((.(((.(--..(((((((.(((.....))).)))))))..)...)))))))))). ( -24.90, z-score = -2.79, R) >droEre2.scaffold_4644 473632 69 + 2521924 CCGCUGCACGAUAACCGCAAACCGCAAAUCGAUAGAUGCGCGCUUUCGAUUGGCUAUGUGUGUUAUCGU .......((((((((((((....((.((((((.((.......)).)))))).))..)))).)))))))) ( -21.60, z-score = -1.38, R) >droYak2.chrX 440109 67 + 21770863 CCGCUGCACGAUAACCGCAAA--UUCCAUCGAUAGCUGCCUGCUUUCGAUAUACUACGCGUGUUAUCGC ........((((((((((...--....(((((.(((.....))).))))).......))).))))))). ( -21.64, z-score = -3.68, R) >droSec1.super_10 365572 67 + 3036183 CAGCUGCACAAUAACCAUGAG--AACAAUCGAUAGCUUCGCGCUUUCGAUUAGCUGUGUGUGUUAUCGG ........(.(((((((((((--...((((((.(((.....))).))))))..)).)))).))))).). ( -15.60, z-score = -0.29, R) >droSim1.chrX 402514 67 + 17042790 CCGCUGCACAAUAACCAUGAG--AACAAUCGAUAGCUUCGCGCUUUCGAUUAGCUGUGUGUGUUAUCGG (((..(((((...((....((--...((((((.(((.....))).))))))..)))).)))))...))) ( -17.70, z-score = -1.19, R) >consensus CCGCUGCACGAUAACCACGAG__AACAAUCGAUAGCUUCGCGCUUUCGAUUAGCUGUGUGUGUUAUCGG ........(((((((((((.......((((((.(((.....))).)))))).....)))).))))))). (-16.40 = -16.96 + 0.56)
| Location | 519,222 – 519,289 |
|---|---|
| Length | 67 |
| Sequences | 5 |
| Columns | 69 |
| Reading direction | reverse |
| Mean pairwise identity | 82.15 |
| Shannon entropy | 0.30933 |
| G+C content | 0.49846 |
| Mean single sequence MFE | -21.22 |
| Consensus MFE | -17.08 |
| Energy contribution | -17.36 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.957692 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
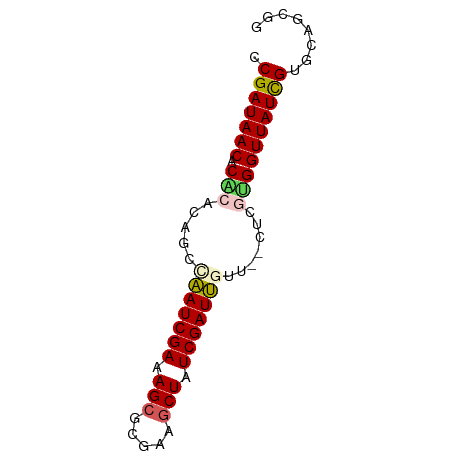
>dm3.chrX 519222 67 - 22422827 CCGAUAACACGCACAGCCAAUCGAAAGCGCGAAGCUAUCGAUUGUU--CUCGUGGUUAUCGUGCAGCGG .(((((((.(((.....(((((((.(((.....))).)))))))..--...))))))))))........ ( -23.80, z-score = -2.43, R) >droEre2.scaffold_4644 473632 69 - 2521924 ACGAUAACACACAUAGCCAAUCGAAAGCGCGCAUCUAUCGAUUUGCGGUUUGCGGUUAUCGUGCAGCGG ((((((((.(.((.((((...(....).(((.(((....))).))))))))).)))))))))....... ( -20.10, z-score = -0.97, R) >droYak2.chrX 440109 67 - 21770863 GCGAUAACACGCGUAGUAUAUCGAAAGCAGGCAGCUAUCGAUGGAA--UUUGCGGUUAUCGUGCAGCGG ((((((((.((((.....((((((.(((.....))).))))))...--..))))))))))))....... ( -26.60, z-score = -3.66, R) >droSec1.super_10 365572 67 - 3036183 CCGAUAACACACACAGCUAAUCGAAAGCGCGAAGCUAUCGAUUGUU--CUCAUGGUUAUUGUGCAGCUG .(((((((......(((.((((((.(((.....))).)))))))))--......)))))))........ ( -17.80, z-score = -1.58, R) >droSim1.chrX 402514 67 - 17042790 CCGAUAACACACACAGCUAAUCGAAAGCGCGAAGCUAUCGAUUGUU--CUCAUGGUUAUUGUGCAGCGG .(((((((......(((.((((((.(((.....))).)))))))))--......)))))))........ ( -17.80, z-score = -1.43, R) >consensus CCGAUAACACACACAGCCAAUCGAAAGCGCGAAGCUAUCGAUUGUU__CUCGUGGUUAUCGUGCAGCGG .(((((((.(((.....(((((((.(((.....))).))))))).......))))))))))........ (-17.08 = -17.36 + 0.28)

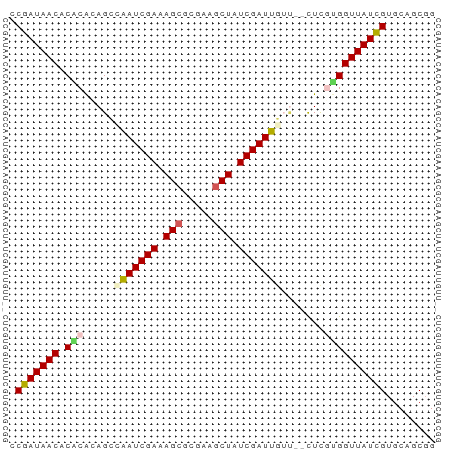
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:03:12 2011