
| Sequence ID | dm3.chrX |
|---|---|
| Location | 513,185 – 513,278 |
| Length | 93 |
| Max. P | 0.629083 |

| Location | 513,185 – 513,278 |
|---|---|
| Length | 93 |
| Sequences | 7 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 76.59 |
| Shannon entropy | 0.45538 |
| G+C content | 0.34939 |
| Mean single sequence MFE | -18.10 |
| Consensus MFE | -8.14 |
| Energy contribution | -8.47 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.629083 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
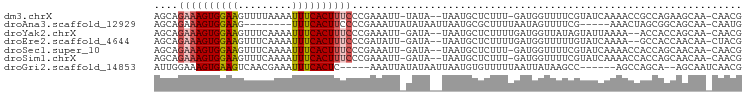
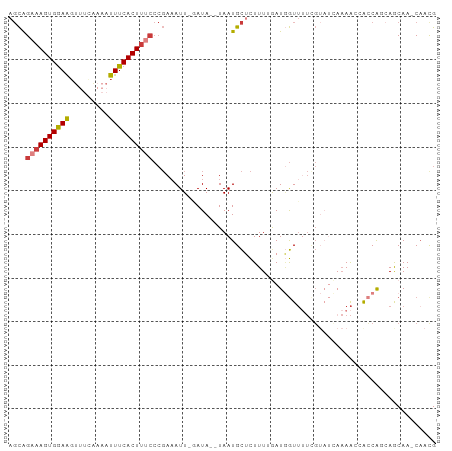
>dm3.chrX 513185 93 + 22422827 AGCAGAAAGUGGAAGUUUUAAAAUUUCACUUUCCCGAAAUU-UAUA--UAAUGCUCUUU-GAUGGUUUUCGUAUCAAAACCGCCAGAAGCAA-CAACG ....((((((((((.........))))))))))........-....--...((((.(((-(.(((((((......))))))).)))))))).-..... ( -22.50, z-score = -3.09, R) >droAna3.scaffold_12929 418460 84 + 3277472 AGCAGAAAGUGGAAG--------UUUCACUUCCCCGAAAUUAUAUAAUUAAUGCGCUUUUAAUAGUUUUCG-----AAACUAGCGGCAGCAA-CAAUG .((.....(.(((((--------.....))))).)...((((......))))))(((.((..(((((....-----.)))))..)).)))..-..... ( -14.30, z-score = -0.29, R) >droYak2.chrX 434096 92 + 21770863 AGCAGAAAGUGGAAGUUUCAAAAUUUCACUUUCCCGAAAUU-GAUA--UAAUGCUCUUUUGAUGGUUAUAGUAUUAAAA--ACCACCAGCAA-CAACG .((.((((((((((.........))))))))))......((-((((--(.(((..(.......)..))).)))))))..--.......))..-..... ( -16.60, z-score = -1.41, R) >droEre2.scaffold_4644 467633 92 + 2521924 AGCAGAAAGUGGAAGUUUCAAAAUUUCACUUUCCCGAUAUU-GAUA--UAAUGCUCUUUUGAUGGUUUUUGUAUCAAAA--GCCACCAACAA-CUACG ((((((((((((((.........))))))))))....(((.-...)--)).))))...(((.(((((((((...)))))--)))).)))...-..... ( -21.10, z-score = -2.69, R) >droSec1.super_10 359745 93 + 3036183 AGCAGAAAGUGGAAGUUUCAAAAUUUCACUUUCCCGAAAUU-GAUA--UAAUGCUCUUU-GAUGGUUUUCGUAUCAAAACCACCAGCAACAA-CAACG .((.((((((((((.........))))))))))........-....--..........(-(.(((((((......))))))).)))).....-..... ( -19.10, z-score = -1.95, R) >droSim1.chrX 396277 93 + 17042790 AGCAGAAAGUGGAAGUUUCAAAAUUUCACUUUCCCGAAAUU-GAUA--UAAUGCUCUUU-GAUGGUUUUCGUAUCAAAACCACCAGCAACAA-CAACG .((.((((((((((.........))))))))))........-....--..........(-(.(((((((......))))))).)))).....-..... ( -19.10, z-score = -1.95, R) >droGri2.scaffold_14853 7099916 85 - 10151454 AUUGGAAAGUGAAGUCAACGAAAUUUCACUC-----AAAUUAUAUAAUUAAUGUGUUUUUAAUUAUAAGCC------AGCCAGCA--AGCAAUCAACG ..(((..((((((((.......)))))))).-----......(((((((((.......)))))))))..))------)((.....--.))........ ( -14.00, z-score = -1.58, R) >consensus AGCAGAAAGUGGAAGUUUCAAAAUUUCACUUUCCCGAAAUU_GAUA__UAAUGCUCUUUUGAUGGUUUUCGUAUCAAAACCACCAGCAGCAA_CAACG ....((((((((((.........))))))))))................................................................. ( -8.14 = -8.47 + 0.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:03:09 2011