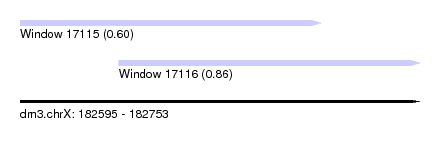

| Sequence ID | dm3.chrX |
|---|---|
| Location | 182,595 – 182,753 |
| Length | 158 |
| Max. P | 0.859317 |

| Location | 182,595 – 182,714 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.76 |
| Shannon entropy | 0.33059 |
| G+C content | 0.36986 |
| Mean single sequence MFE | -27.84 |
| Consensus MFE | -18.78 |
| Energy contribution | -18.58 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.598623 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 182595 119 + 22422827 UAAUGACGCAGAUGGGAGAC-GUUUCUGUAAAAUCAGACGAUAUGUAUUAAAGAACCGGAUUCGGAAAUCUGAAUCCAGGCCAAAAGAUCAAAUAUCGCUCUACGAAAUUAUUGUAUUAA ((((((.(((((..(....)-...)))))......(((((((((...........((((((((((....)))))))).))............)))))).)))......))))))...... ( -28.40, z-score = -2.04, R) >droSim1.chrX 126613 114 + 17042790 UAAUGACGAAGAUGGGGGAC-UUUUCUGUAAAAUCAGCCGAUUUGUAUUAAGGAAUCGGAUUCGGAAAACUGAAUCCAGGUCAAAAGAUCAAAUAUCGCUCUGCGAAAUUAUUAC----- ((((((.(((((.((....)-)..))).........((((((((.(....).)))))((((((((....)))))))).))).......)).....((((...))))..)))))).----- ( -26.10, z-score = -1.18, R) >droSec1.super_10 57143 114 + 3036183 UAAUGACGAAAAUGGGGGAC-UUUUCUGUAAAAUCAGACGAUUUGUAUUAAGGAAUCGGAUUCGGAAAACUGAAUCCAGGUCAAAAGAUCAAAUAUCGCUCUGCGAAAUUAUUAC----- ((((((.((((..((....)-)))))........((((((((...............((((((((....)))))))).((((....))))....)))).)))).....)))))).----- ( -25.90, z-score = -1.76, R) >droYak2.chrX 125190 114 + 21770863 CAAUGAUAAAGAUGGGGGGA-AUUUCUGUGAAAUCAGCCGAUAUGUAUUAAGGAAUCGGAUUCGGAAAUAUGAAUCCAGAUCAAGUGAUCAACUAUCGCUUCGUGAUUUAGGUUU----- ...(((((...((.((..((-.((((...))))))..)).))...)))))..((((((((((((......)))))))((((((((((((.....))))))...)))))).)))))----- ( -26.70, z-score = -0.61, R) >droEre2.scaffold_4644 130001 115 + 2521924 UAAUGAUGAAGAUGGAAGGGAGUUUCUGUGAAAGCAGCCGAUUUGUAAUGAUGAAUCGGAUUCGCACAUGAGAAUCCAGAUCAAAUGAUCAACUAUCGCUCUGUGAUAUUAUUAA----- ((((((((..(((((...(((.((((((((...(...((((((..(....)..))))))...).)))).)))).))).((((....))))..))))).(.....).)))))))).----- ( -32.10, z-score = -2.26, R) >consensus UAAUGACGAAGAUGGGGGAC_GUUUCUGUAAAAUCAGCCGAUUUGUAUUAAGGAAUCGGAUUCGGAAAUCUGAAUCCAGGUCAAAAGAUCAAAUAUCGCUCUGCGAAAUUAUUAC_____ ..............((((.......(((......))).((((...............((((((((....)))))))).((((....))))....)))))))).................. (-18.78 = -18.58 + -0.20)


| Location | 182,634 – 182,753 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.66 |
| Shannon entropy | 0.40677 |
| G+C content | 0.37826 |
| Mean single sequence MFE | -27.59 |
| Consensus MFE | -14.20 |
| Energy contribution | -14.28 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.859317 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 182634 119 + 22422827 AUAUGUAUUAAAGAACCGGAUUCGGAAAUCUGAAUCCAGGCCAAAAGAUCAAAUAUCGCUCUACGAAAUUAUUGUAUUAAACAGAAACUAAAUCAAUGCUGCUG-GGACCGGUAGCAGCC .................((((((((....)))))))).(((.....(((..(((.(((.....))).))).((((.....)))).......)))..((((((((-....))))))))))) ( -30.10, z-score = -2.62, R) >droSim1.chrX 126652 113 + 17042790 AUUUGUAUUAAGGAAUCGGAUUCGGAAAACUGAAUCCAGGUCAAAAGAUCAAAUAUCGCUCUGCGAAAUUAUUACA-----CAGAAACUAAAUCUAUGCUGCUG-GGACCGGUA-CAGUC ..(((((((........((((((((....)))))))).((((...((....(((.((((...)))).)))....((-----.(((.......))).))...)).-.))))))))-))).. ( -26.70, z-score = -1.88, R) >droSec1.super_10 57182 113 + 3036183 AUUUGUAUUAAGGAAUCGGAUUCGGAAAACUGAAUCCAGGUCAAAAGAUCAAAUAUCGCUCUGCGAAAUUAUUACA-----CAGAAACUAAAUCAAUGCUGCUG-GGACCGGUA-CAGUC ..(((((((........((((((((....)))))))).((((.........(((.((((...)))).)))....((-----(((..............))).))-.))))))))-))).. ( -26.54, z-score = -1.88, R) >droYak2.chrX 125229 104 + 21770863 AUAUGUAUUAAGGAAUCGGAUUCGGAAAUAUGAAUCCAGAUCAAGUGAUCAACUAUCGCUUCGUGAUUUAGGUUUU-----UUAAAAUAAUUUCAAAGCUUUUG-GGAUC---------- ...(((.(((((((((((((((((......)))))))((((((((((((.....))))))...)))))).))))))-----)))).)))((..(((.....)))-..)).---------- ( -26.80, z-score = -2.80, R) >droEre2.scaffold_4644 130041 115 + 2521924 AUUUGUAAUGAUGAAUCGGAUUCGCACAUGAGAAUCCAGAUCAAAUGAUCAACUAUCGCUCUGUGAUAUUAUUAAA-----ACGAAACUAAAUCAAAGCUCCUGCGGUCCGGUCGCAGCC .(((((..((((((...((((((........)))))).((((....))))...((((((...))))))))))))..-----)))))...............((((((.....)))))).. ( -27.80, z-score = -1.61, R) >consensus AUUUGUAUUAAGGAAUCGGAUUCGGAAAUCUGAAUCCAGGUCAAAAGAUCAAAUAUCGCUCUGCGAAAUUAUUACA_____CAGAAACUAAAUCAAUGCUGCUG_GGACCGGUA_CAGCC .................((((((((....)))))))).((((....)))).....((((...))))...................................................... (-14.20 = -14.28 + 0.08)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:02:28 2011