
| Sequence ID | dm3.chrX |
|---|---|
| Location | 44,390 – 44,452 |
| Length | 62 |
| Max. P | 0.716459 |

| Location | 44,390 – 44,452 |
|---|---|
| Length | 62 |
| Sequences | 5 |
| Columns | 65 |
| Reading direction | forward |
| Mean pairwise identity | 59.34 |
| Shannon entropy | 0.73092 |
| G+C content | 0.34617 |
| Mean single sequence MFE | -9.42 |
| Consensus MFE | -4.50 |
| Energy contribution | -4.66 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 2.00 |
| Mean z-score | -0.57 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.716459 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
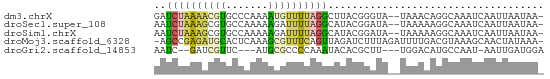
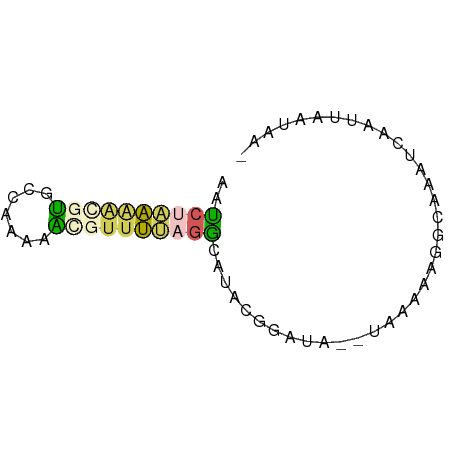
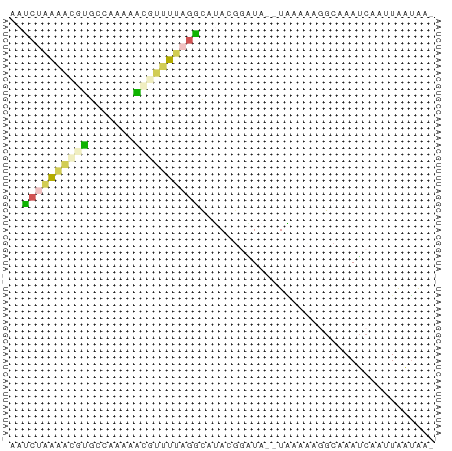
>dm3.chrX 44390 62 + 22422827 GAUCUAAAACGUGCCCAAAAUGUUUUAGGCUUACGGGUA--UAAACAGGCAAAUCAAUUAAUAA- ((((((((((((.......)))))))))((((.......--.....))))..))).........- ( -9.90, z-score = -0.82, R) >droSec1.super_108 34233 62 + 76128 AAUCUAAAGCGUGCCAAAAAGAUUUUAGGCAUACGGAUA--UAAAAAGGCAAAUCAAUUAAUAA- .((((...(..((((.(((....))).))))..))))).--.......................- ( -7.10, z-score = -0.32, R) >droSim1.chrX 28590 62 + 17042790 AAUCUAAAGCGUGCCAAAAAGAUUUUAGGCAUACGGAUA--UAAAAAGGCAAAUCAAUUAAUAA- .((((...(..((((.(((....))).))))..))))).--.......................- ( -7.10, z-score = -0.32, R) >droMoj3.scaffold_6328 1410791 63 + 4453435 -AGCCGAGAUGCACUCAAAGCGUUUCAGUUAGAUCUUUAGAUUUUGACGUAAAGCAACUAUAAA- -.((.(((((((.......))))))).((((((((....)).)))))).....)).........- ( -13.90, z-score = -1.77, R) >droGri2.scaffold_14853 5638132 56 - 10151454 AAUC--GAUCGUUC---AUGCGCCCCAAAUACACGCUU---UGGACAUGCCAAU-AAUUGAUGGA .(((--(((..(((---(((....(((((.......))---))).))))..)).-.))))))... ( -9.10, z-score = 0.38, R) >consensus AAUCUAAAACGUGCCAAAAACGUUUUAGGCAUACGGAUA__UAAAAAGGCAAAUCAAUUAAUAA_ ..((((((((((.......)))))))))).................................... ( -4.50 = -4.66 + 0.16)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:02:21 2011