
| Sequence ID | dm3.chrXHet |
|---|---|
| Location | 160,605 – 160,738 |
| Length | 133 |
| Max. P | 0.975541 |

| Location | 160,605 – 160,738 |
|---|---|
| Length | 133 |
| Sequences | 3 |
| Columns | 133 |
| Reading direction | forward |
| Mean pairwise identity | 61.71 |
| Shannon entropy | 0.55283 |
| G+C content | 0.58241 |
| Mean single sequence MFE | -54.40 |
| Consensus MFE | -19.56 |
| Energy contribution | -23.23 |
| Covariance contribution | 3.68 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.728871 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
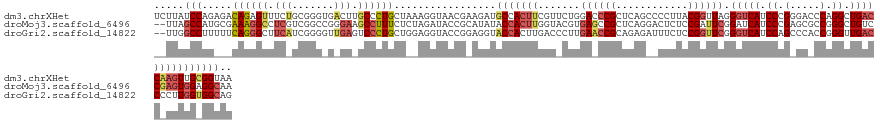
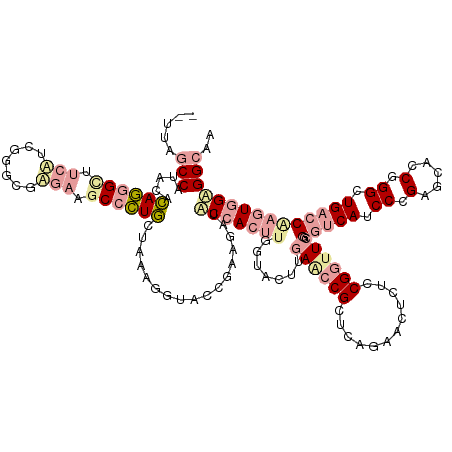
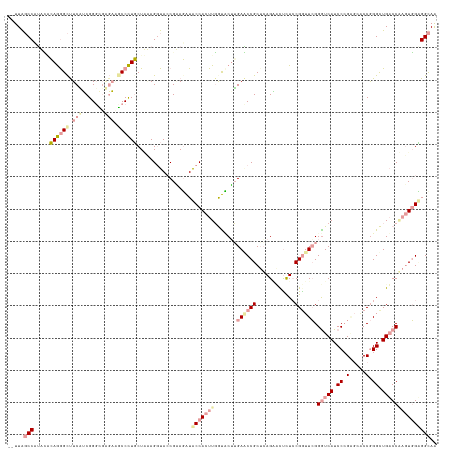
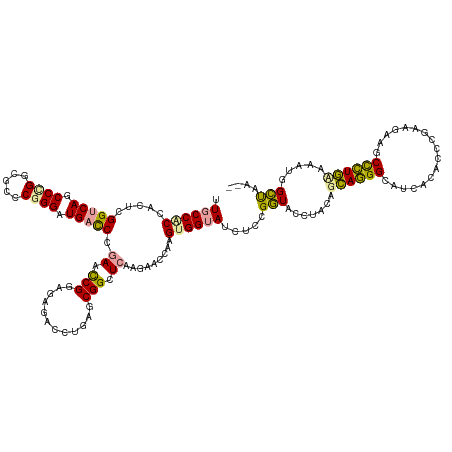
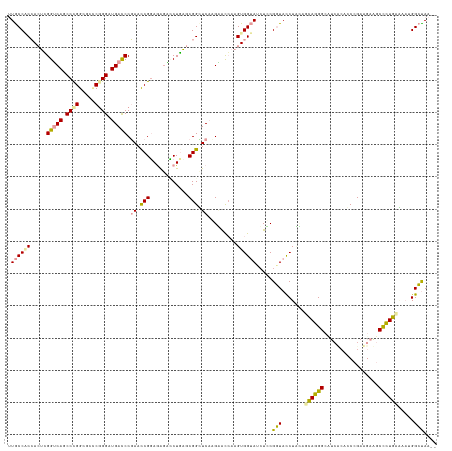
>dm3.chrXHet 160605 133 + 204112 UCUUAUCCAGAGACAGAGUUUCUGCGGGUGACUUGCCCUGCUAAAGGUAACGAAGAUGCCACUUCGUUCUGGACCCGCUCAGCCCCUUACGGUUAGGGUCAUCCCGGGACCCAGGCUGACCAAGUUGCGGUAA .....(((((((.(..(((....((((((.....)))).))....((((.......)))))))..)))))))).((((((((((((....))...(((((.......))))).)))))).......))))... ( -45.61, z-score = -0.56, R) >droMoj3.scaffold_6496 1647639 131 + 26866924 --UUAGCCAUGCGAAAGGCCUCGUCGGCCGGGAAGCCUUUCUCUAGAUACCGCAUAUACCACUUGGUACGUGAGCCGCUCAGGACUCUCCGAUUCGGAUCAUCCCGAGCGCCGGGCUGUCCGAGUGGAGGCAA --...((((((((...((((.....))))(((((....))))).......)))))...((((((((.((((..(.(((((.(((...((((...))))...))).))))).)..)).)))))))))).))).. ( -55.50, z-score = -2.29, R) >droGri2.scaffold_14822 968639 131 + 1097346 --UUGGCCUUUUUCAGGGCUUCAUCGGGGUUGAGUCCCUGCUGGAGGUACCGGAGGUACCACUUGACCCUUGAACCGCAGAGAUUUCUCCGGUUCGGGUCAUCCAGCCCACCGGGUUGACCCCUUGGUGGCAG --..(((((......)))))((((((((((..(..(((.((((((((((((...))))))...((((((..((((((..(((....))))))))))))))))))))).....))))..)))))..)))))... ( -62.10, z-score = -3.04, R) >consensus __UUAGCCAUACACAGGGCUUCAUCGGGCGAGAAGCCCUGCUAAAGGUACCGAAGAUACCACUUGGUACUUGAACCGCUCAGAACUCUCCGGUUCGGGUCAUCCCGAGCACCGGGCUGACCAAGUGGAGGCAA .....(((.....((((((.(((.......))).)))))).................(((((((.......((((((............)))))).(((((.((.(.....).)).))))))))))))))).. (-19.56 = -23.23 + 3.68)
| Location | 160,605 – 160,738 |
|---|---|
| Length | 133 |
| Sequences | 3 |
| Columns | 133 |
| Reading direction | reverse |
| Mean pairwise identity | 61.71 |
| Shannon entropy | 0.55283 |
| G+C content | 0.58241 |
| Mean single sequence MFE | -55.94 |
| Consensus MFE | -24.30 |
| Energy contribution | -23.76 |
| Covariance contribution | -0.54 |
| Combinations/Pair | 1.34 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.93 |
| SVM RNA-class probability | 0.975541 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
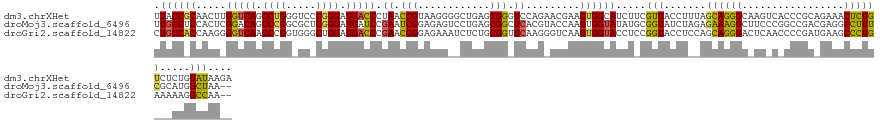
>dm3.chrXHet 160605 133 - 204112 UUACCGCAACUUGGUCAGCCUGGGUCCCGGGAUGACCCUAACCGUAAGGGGCUGAGCGGGUCCAGAACGAAGUGGCAUCUUCGUUACCUUUAGCAGGGCAAGUCACCCGCAGAAACUCUGUCUCUGGAUAAGA ...((((.......((((((.(((((.......)))))...((....))))))))))))(((((((((((((......))))))...........(((.......)))((((.....)))).))))))).... ( -49.21, z-score = -1.05, R) >droMoj3.scaffold_6496 1647639 131 - 26866924 UUGCCUCCACUCGGACAGCCCGGCGCUCGGGAUGAUCCGAAUCGGAGAGUCCUGAGCGGCUCACGUACCAAGUGGUAUAUGCGGUAUCUAGAGAAAGGCUUCCCGGCCGACGAGGCCUUUCGCAUGGCUAA-- ..((((((....)))..(((((.((((((((((..((((...))))..)))))))))).).((..((((....))))..)).))........((((((((((.........))))))))))))..)))...-- ( -56.00, z-score = -2.11, R) >droGri2.scaffold_14822 968639 131 - 1097346 CUGCCACCAAGGGGUCAACCCGGUGGGCUGGAUGACCCGAACCGGAGAAAUCUCUGCGGUUCAAGGGUCAAGUGGUACCUCCGGUACCUCCAGCAGGGACUCAACCCCGAUGAAGCCCUGAAAAAGGCCAA-- ..(((.(((..(((....)))..)))((((((((((((((((((.(((....))).))))))..))))))...((((((...))))))))))))((((..(((.......)))..))))......)))...-- ( -62.60, z-score = -3.78, R) >consensus UUGCCACCACUCGGUCAGCCCGGCGCCCGGGAUGACCCGAACCGGAGAGACCUGAGCGGCUCAAGAACCAAGUGGUAUCUCCGGUACCUACAGCAGGGCAUCACACCCGAAGAAGCCCUGAAAAUGGCUAA__ .((((((.....(((((.((((.....)))).))))).((.(((............))).)).........)))))).....(((.......((((((.................)))))).....))).... (-24.30 = -23.76 + -0.54)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:02:08 2011