
| Sequence ID | dm3.chr4 |
|---|---|
| Location | 746,397 – 746,517 |
| Length | 120 |
| Max. P | 0.588749 |

| Location | 746,397 – 746,517 |
|---|---|
| Length | 120 |
| Sequences | 15 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.46 |
| Shannon entropy | 0.51936 |
| G+C content | 0.43056 |
| Mean single sequence MFE | -32.27 |
| Consensus MFE | -11.87 |
| Energy contribution | -11.37 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.57 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.588749 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
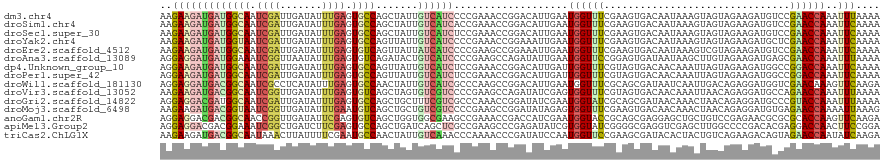
>dm3.chr4 746397 120 + 1351857 AAGAAGAUGAUGGCAAUCGAUUGAUAUUUGAGUGCCAGCUAUUGUCAUCCCCGAAACCGGACAUUGAAUGGUUUCGAAGUGACAAUAAAGUAGUAGAAGAUGUCCGAACCAAAUUUAAAA .....(((..(((((.((((.......)))).)))))..(((((((((...((((((((.........))))))))..)))))))))..............)))................ ( -29.60, z-score = -2.85, R) >droSim1.chr4 580324 120 + 949497 AAGAAGAUGAUGGCAAUCGAUUGAUAUUUGAGUGCCAGCUAUUGUCAUCACCGAAACCGGACAUUGAAUGGUUUCGAAGUGACAAUAAAGUAGUAGAAGAUGUCCGAACCAAAUUCAAAA ..(((..((.(((((.((((.......)))).)))))..(((((((((...((((((((.........))))))))..)))))))))......................))..))).... ( -31.60, z-score = -3.00, R) >droSec1.super_30 463605 120 + 664411 AAGAAGAUGAUGGCAAUCGAUUGAUAUUUGAGUGCCAGCUAUUGUCAUCUCCGAAACCGGACAUUGAAUGGUUUCGAAGUGACAAUAAAGUAGUAGAAGAUGUCCGAACCAAAUUCAAAA ..(((..((.(((((.((((.......)))).)))))..((((((((.((.((((((((.........)))))))).))))))))))......................))..))).... ( -32.60, z-score = -3.07, R) >droYak2.chr4 810313 120 + 1374474 AAGAAGAUGAUGGUAAUCGAUUGAUAUUUGAGUGCCAGUUAUUGUCAUCCCCGAAACCGGAAAUUGAAUGGUUUCGAAGUGACAAUAAAGUAGUAGAAGAUGCUCGAACCAAAUUCAAAA ..(((..((.(((((.((((.......)))).))))).((((((((((...((((((((.........))))))))..)))))))))).(.((((.....)))))....))..))).... ( -31.80, z-score = -3.74, R) >droEre2.scaffold_4512 769179 120 - 1286254 AAGAAGAUGAUGGCAAUCGAUUGAUAUUUGAGUGUCAGUUAUUAUCAUCCCCGAAGCCGGAAAUUGAAUGGUUUCGAAGUGACAAUAAAGUCGUAGAAGAUGUCCGAACCAAAUUCAAAA ........((((..(((.(((((((((....))))))))))))..)))).(((....)))...(((((((((((.((..((((......)))).........)).))))))..))))).. ( -26.20, z-score = -1.29, R) >droAna3.scaffold_13089 573831 120 + 978470 AGGAGGAUGAUGGAAAUCGGUUAAUAUUUGAGUGUCAGAUACUGUCAUCCCCGAAGCCAGAUAUUGAAUGGUUCCGGAGUGAUAAUAAGCUUGUAGAAGAUGAGCGAACCAAAUUUAAAA .((.(((((((((...........((((((.....)))))))))))))))(((.(((((.........))))).)))...........(((..(.....)..)))...)).......... ( -28.80, z-score = -1.68, R) >dp4.Unknown_group_10 47592 120 + 72859 AGGAAGAUGAUGGCAAUCGAUUGAUAUUUGAGUGCCAGUUAUUGUCAUCUCCGAAACCGGACAUUGAUUGGUUUCGUAGUGACAACAAAUUAGUAGAAGAUGGCCGGACCAAAUUCAAAA .....((((((((((.((((.......)))).))))).)))))((((.((.(((((((((.......))))))))).))))))............(((..(((.....)))..))).... ( -32.30, z-score = -1.85, R) >droPer1.super_42 549830 120 + 623919 AGGAAGAUGAUGGCAAUCGAUUGAUAUUUGAGUGCCAGUUAUUGUCAUCUCCGAAACCGGACAUUGAUUGGUUUCGUAGUGACAACAAAUUAGUAGAAGAUGGCCGGACCAAAUUCAAAA .....((((((((((.((((.......)))).))))).)))))((((.((.(((((((((.......))))))))).))))))............(((..(((.....)))..))).... ( -32.30, z-score = -1.85, R) >droWil1.scaffold_181130 1667194 120 + 16660200 AGGAGGAUGACGGCAAUCGCCUCAUAUUUGAGUGCCAACUAUUGUCAUCGCCCAAGCCGGACAUUGAAUGGUUUCGCAGCGAUAAUCAAUUGACAGAGGAUGGUCGAACAAAGUUCAAGA .((..((((((((.....((((((....)))).))......))))))))..)).....((((.(((..((((((((...))).))))).(((((........))))).))).)))).... ( -30.00, z-score = 0.18, R) >droVir3.scaffold_13052 1536884 120 + 2019633 AAGAAGAUGACGGCAAUCGGUUGAUAUUUGAGUGUCAGCUAGUGUCGUCCCCGAAGCCAGAUAUCGAGUGGUUUCGUAGUGACAACAAAUUAACAGAGGAUGCCAGAACCAAAUUUAAAA ...........((((.(((((((((((....)))))))))..(((((.(..((((((((.........))))))))..)))))).............)).))))................ ( -32.20, z-score = -2.54, R) >droGri2.scaffold_14822 542964 120 + 1097346 AGGAGGACGAUGGCAAUCGAUUGAUAUUUGAGUGCCAGCUGCUUUCGUCGCCCAAACCGGAUAUCGAAUGGUAUCGCAGCGAUAACAAACUAACAGAGGAUGCCCGUACCAAAUUUAAAA ....(((((.(((((.((((.......)))).)))))(((((......((.......))((((((....)))))))))))........................))).)).......... ( -28.60, z-score = -0.17, R) >droMoj3.scaffold_6498 539892 120 + 3408170 AAGAAGAUGACGGUAAUCGGUUGAUAUUUGAAUGUCAGCUGCUGUCGUCCCCGAAGCCGGAUAUAGAGUGGUUUCGAAGUGACAACAAACUAACAGAGGAUGUGAGAACCAAAAUUAAAG .....(((((((((....(((((((((....))))))))))))))))))..((((((((.........))))))))..((....(((..((.....))..)))....))........... ( -31.90, z-score = -2.41, R) >anoGam1.chr2R 55541216 120 - 62725911 AGGAGGACGACGGCAACCGGUUGAUAUUCGAGUGUCAGCUGGUGGCGAAGCCGAAACCGACCAUCGAAUGGUACCGCAGCGAGGAGCUGCUGUCCGAGAACGCGCGCACCAAGUUCAAGA ....(((((.(((..((((((((((((....))))))))))))(((...)))....)))(((((...)))))...(((((.....))))))))))..((((...........)))).... ( -44.00, z-score = -1.20, R) >apiMel3.Group2 14086011 120 - 14465785 AGGAGGACGACGGAAAUCGGCUGAUCUUCGAGUGCCAGCUGAUCAGCUCGCCGAAGCCCGAGAUAUCGUGGUAUCGGGGCGAGGUCGAGCUUGGCCCCGACACGAGGACCAACUUCCGGA .(((((.(((......)))..((.((((((.(.((.((((....)))).)))...((((((....))).))).(((((((.(((.....))).)))))))..)))))).)).)))))... ( -48.20, z-score = -0.68, R) >triCas2.ChLG1X 1228485 120 + 8109244 AAGAAGAUGACGGCAAUAAACUUAUUUUCGAAUGCCAACUAUUGUCAAACCCAAAACCCGAUAUCCAAUGGUUCCGAAGCGAUACACUACUGUCAGAAGACAGUAGAACCAAUAUCAAGA .....((((..((............(((((...((((.....((((.............)))).....))))..))))).......((((((((....))))))))..))..)))).... ( -23.92, z-score = -2.38, R) >consensus AAGAAGAUGAUGGCAAUCGAUUGAUAUUUGAGUGCCAGCUAUUGUCAUCCCCGAAACCGGACAUUGAAUGGUUUCGAAGUGACAAUAAACUAGUAGAAGAUGCCCGAACCAAAUUCAAAA ..(((((((((((((.((((.......)))).)))).......))))))...................((((((...............................))))))..))).... (-11.87 = -11.37 + -0.50)

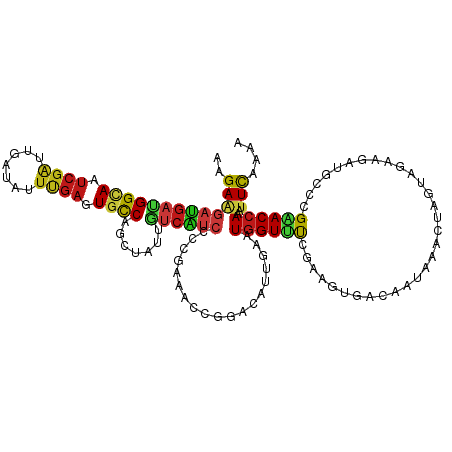
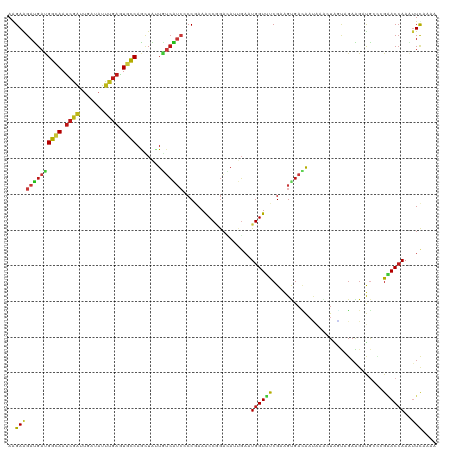
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:01:31 2011