
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 27,533,771 – 27,533,866 |
| Length | 95 |
| Max. P | 0.999707 |

| Location | 27,533,771 – 27,533,866 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 61.33 |
| Shannon entropy | 0.64802 |
| G+C content | 0.59407 |
| Mean single sequence MFE | -42.56 |
| Consensus MFE | -25.05 |
| Energy contribution | -25.21 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.77 |
| SVM RNA-class probability | 0.995160 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
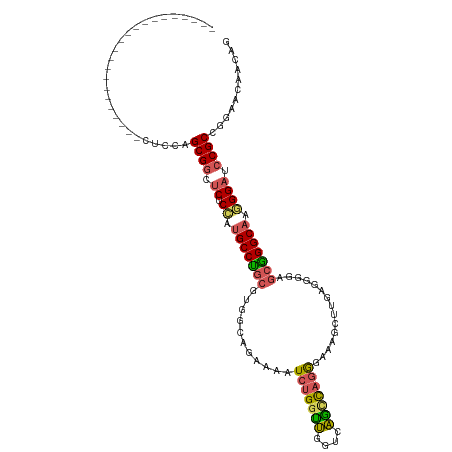
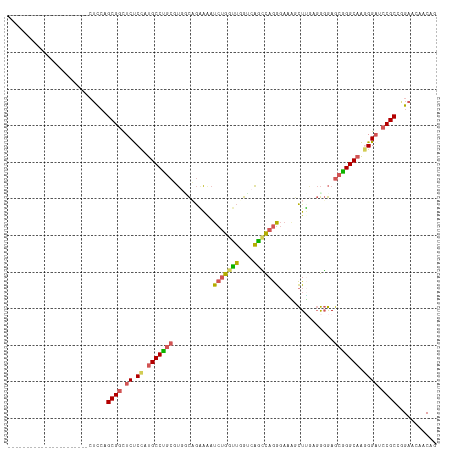
>dm3.chr3R 27533771 95 + 27905053 ----------------------CUCCAGCGCCUCUCCAUGCCUGCGUGGCAGAAGAUCUGGUUGGUCAGCCAGGGAAAGCUUGAGGGGAGCGGGCCAGGGAUCCGCUAGAACAACAG ----------------------(((.(((.((((((((((....))))).)))....((((((....))))))))...))).)))...(((((((....).)))))).......... ( -35.00, z-score = 0.28, R) >droSim1.chr3R 27148752 95 + 27517382 ----------------------CUCCAGCGGCUCUCCAUGCCUGCGUGUCCGAAAAUCUGGUUGGUCAGCCAGGGAAAGCUUGAGGGGAGCGGGCAAGGGAUCCGCCAGAACAACAG ----------------------.((..((((.((.((.(((((((.(.(((.....(((((((....)))))))..........))).)))))))).)))).))))..))....... ( -36.76, z-score = -0.75, R) >droSec1.super_31 188200 95 + 568655 ----------------------CUCCAGCGGCUCUCCAUGCCUGCGUGGCCGAAAAUCUGGUUGGUCAGUCAGGGAGAGCUUGAGGGGAGCGGGCAACGGAUCCGCCGGAACAACAG ----------------------.(((.((((.(((...(((((((.(((((((........))))))).(((((.....))))).....)))))))..))).)))).)))....... ( -38.30, z-score = -0.80, R) >droYak2.chr3R 28467141 105 + 28832112 ------------UGCUCUGCUGCUCCAGCGGCUCUCCAUGCCCACUUGCCAGCAAAUCUGGCUCGGAAGCCAGGGAUUCCUGAUGGGGAGCGGGCAAGGGCUCCGCUGGAACAACAG ------------....(((.((.((((((((...(((.(((((.(((.(((.((..(((((((....)))))))......)).))).))).))))).)))..)))))))).)).))) ( -51.70, z-score = -2.67, R) >droAna3.scaffold_12911 1805300 117 - 5364042 CUUCUUGUGGAUAUUCUGGAAGGAUAGGCGGUUCCCUCUGCCUGCUCAUUAAGGAAUCCGAUCUAGGGAGUAAAUCGGAUUCCUCAUAAACAGGCAGAGGAUCCGCAGGGAUUUCAU .(((((((((((((((.....)))))........((((((((((.......(((((((((((.((.....)).)))))))))))......)))))))))).))))))))))...... ( -51.02, z-score = -4.35, R) >consensus ______________________CUCCAGCGGCUCUCCAUGCCUGCGUGGCAGAAAAUCUGGUUGGUCAGCCAGGGAAAGCUUGAGGGGAGCGGGCAAGGGAUCCGCCGGAACAACAG ...........................((((.((.((.(((((((...........(((((((....)))))))...............))))))).)))).))))........... (-25.05 = -25.21 + 0.16)


| Location | 27,533,771 – 27,533,866 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 61.33 |
| Shannon entropy | 0.64802 |
| G+C content | 0.59407 |
| Mean single sequence MFE | -42.54 |
| Consensus MFE | -22.12 |
| Energy contribution | -22.52 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.44 |
| Mean z-score | -2.89 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.23 |
| SVM RNA-class probability | 0.999707 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
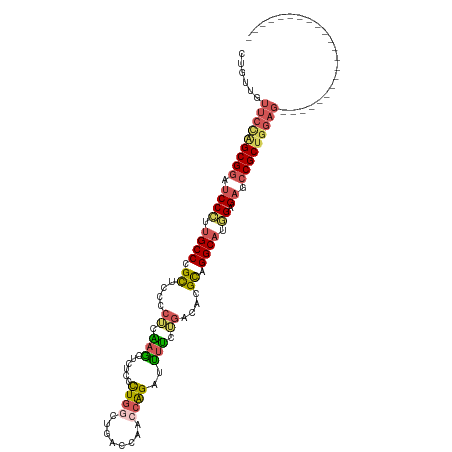
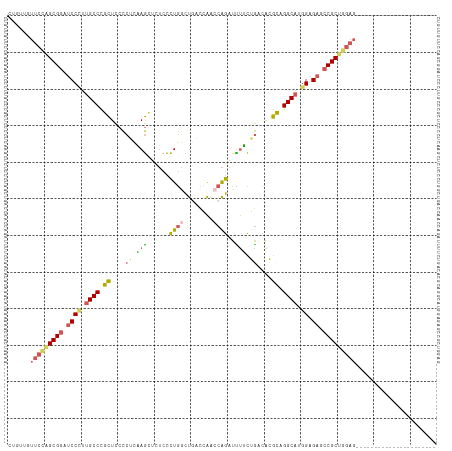
>dm3.chr3R 27533771 95 - 27905053 CUGUUGUUCUAGCGGAUCCCUGGCCCGCUCCCCUCAAGCUUUCCCUGGCUGACCAACCAGAUCUUCUGCCACGCAGGCAUGGAGAGGCGCUGGAG---------------------- .......(((((((.....((((...(((.......)))......(((....))).)))).(((((((((.....)))).)))))..))))))).---------------------- ( -29.50, z-score = 0.81, R) >droSim1.chr3R 27148752 95 - 27517382 CUGUUGUUCUGGCGGAUCCCUUGCCCGCUCCCCUCAAGCUUUCCCUGGCUGACCAACCAGAUUUUCGGACACGCAGGCAUGGAGAGCCGCUGGAG---------------------- .......((..((((.((((.((((.((.....(((.(((......))))))....((........))....)).)))).)).)).))))..)).---------------------- ( -32.50, z-score = -0.64, R) >droSec1.super_31 188200 95 - 568655 CUGUUGUUCCGGCGGAUCCGUUGCCCGCUCCCCUCAAGCUCUCCCUGACUGACCAACCAGAUUUUCGGCCACGCAGGCAUGGAGAGCCGCUGGAG---------------------- .......((((((((.(((((.(((.((...((......((.....))(((......)))......))....)).))))))))...)))))))).---------------------- ( -32.40, z-score = -0.67, R) >droYak2.chr3R 28467141 105 - 28832112 CUGUUGUUCCAGCGGAGCCCUUGCCCGCUCCCCAUCAGGAAUCCCUGGCUUCCGAGCCAGAUUUGCUGGCAAGUGGGCAUGGAGAGCCGCUGGAGCAGCAGAGCA------------ (((((((((((((((..(((.((((((((..(((.((((.....((((((....)))))).)))).)))..)))))))).)).)..)))))))))))))))....------------ ( -63.00, z-score = -5.96, R) >droAna3.scaffold_12911 1805300 117 + 5364042 AUGAAAUCCCUGCGGAUCCUCUGCCUGUUUAUGAGGAAUCCGAUUUACUCCCUAGAUCGGAUUCCUUAAUGAGCAGGCAGAGGGAACCGCCUAUCCUUCCAGAAUAUCCACAAGAAG ...........((((.(((((((((((((((((((((((((((((((.....))))))))))))))).))))))))))))))))..))))........................... ( -55.30, z-score = -8.00, R) >consensus CUGUUGUUCCAGCGGAUCCCUUGCCCGCUCCCCUCAAGCUCUCCCUGGCUGACCAACCAGAUUUUCUGACACGCAGGCAUGGAGAGCCGCUGGAG______________________ .......((((((((.((((.((((.((....((.(((......((((........))))..))).))....)).)))).)).)).))))))))....................... (-22.12 = -22.52 + 0.40)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:00:33 2011