| Sequence ID | dm3.chr3R |
|---|---|
| Location | 27,385,556 – 27,385,647 |
| Length | 91 |
| Max. P | 0.987836 |

| Location | 27,385,556 – 27,385,647 |
|---|---|
| Length | 91 |
| Sequences | 8 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 71.18 |
| Shannon entropy | 0.54656 |
| G+C content | 0.52165 |
| Mean single sequence MFE | -31.38 |
| Consensus MFE | -20.11 |
| Energy contribution | -19.79 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.29 |
| SVM RNA-class probability | 0.987836 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
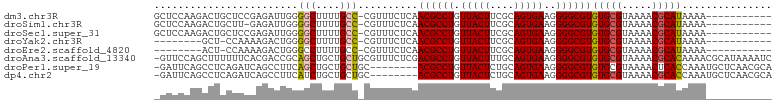
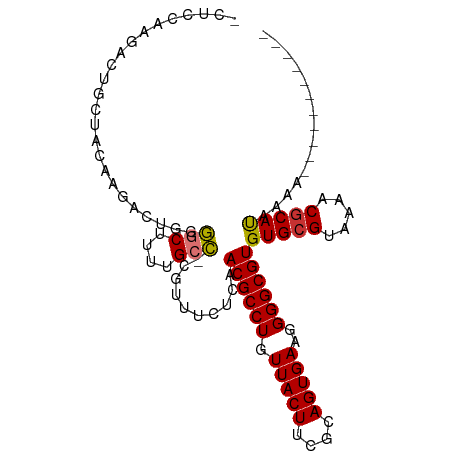
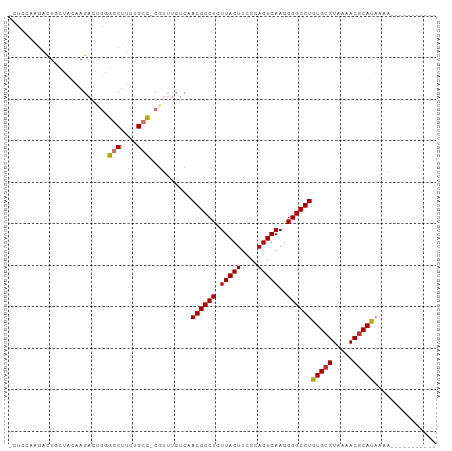
>dm3.chr3R 27385556 91 + 27905053 GCUCCAAGACUGCUCCGAGAUUGGGGCUUUUGCC-CGUUUCUCAACGCCUGUUACUUCGCAGUGAAGGGGCGUGUGCGUAAAACGCAUAAAA----------- (((((...(((((...((((...((((....)))-)...))))(((....))).....)))))...))))).((((((.....))))))...----------- ( -35.50, z-score = -2.81, R) >droSim1.chr3R 27021452 90 + 27517382 GCUCCAAGACUGCUU-GAGAUUGGGGCUUUUGCC-CGUUUCUCAACGCCUGUUACUUCGCAGUGAAGGGGCGUGUGCGUAAAACGCAUAAAA----------- (((((......((((-((((...((((....)))-)...)))))).))((((......))))....))))).((((((.....))))))...----------- ( -35.50, z-score = -3.07, R) >droSec1.super_31 46108 91 + 568655 GCUCCAAGACUGCUCCGAGAUUGGGGCUUUUGCC-CGUUUCUCAACGCCUGUUACUUCGCAGUGAAGGGGCGUGUGCGUAAAACGCAUAAAA----------- (((((...(((((...((((...((((....)))-)...))))(((....))).....)))))...))))).((((((.....))))))...----------- ( -35.50, z-score = -2.81, R) >droYak2.chr3R 28320614 82 + 28832112 --------GCU-CCAAAAGACUGGGGCUUUUGCC-CGUUUCUCAACGCCUGUUACUUCGCAGUGAAGGGGCGUGUGCGUAAAACGCAUAAAA----------- --------(((-((.........((((....)))-).........(((.(((......))))))..))))).((((((.....))))))...----------- ( -27.60, z-score = -1.59, R) >droEre2.scaffold_4820 9879689 82 - 10470090 --------ACU-CCAAAAGACUGGGCCUUUUGCC-CGUUUCUCAACGCCUGUUACUUCGCAGUGAAGGGGCGUGUGCGUAAAACGCAUAAAA----------- --------...-.....((((.((((.....)))-)))))....((((((.(((((....)))))..))))))(((((.....)))))....----------- ( -26.50, z-score = -1.75, R) >droAna3.scaffold_13340 16522555 102 + 23697760 -GUUCCAGCUUUUUUCACGACCGCAGCUGCUGCUGCGUUUCUCGACGCCUGUUACUUUGCAGUGAAGGGGCGUGUGCGUAAAACGCACAAAACGCAUAAAAUC -......((.((((...(((.((((((....))))))....)))((((((.(((((....)))))..))))))(((((.....))))))))).))........ ( -34.90, z-score = -1.83, R) >droPer1.super_19 334694 94 + 1869541 -GAUUCAGCCUCAGAUCAGCCUUCAGCUGCUGCUGC--------ACGCCUGUUACUCUGCAGUGAAGGGGCGUGUGCGUAAAACUCACCAAAUGCUCAACGCA -((((........))))..(((((((((((....))--------).))((((......)))))))))).((((..((((............))))...)))). ( -25.40, z-score = 0.90, R) >dp4.chr2 4635982 94 + 30794189 -GAUUCAGCCUCAGAUCAGCCUUCAUCUGCUGCUGC--------ACGCCUGUUACUCUGCAGUGAAGGGGCGUGUGCGUAAAACGCACCAAAUGCUCAACGCA -....((((..(((((........)))))..)))).--------((((((.(((((....)))))..))))))(((((.....)))))....(((.....))) ( -30.10, z-score = -0.78, R) >consensus _CUCCAAGACUGCUACAAGACUGGGGCUUUUGCC_CGUUUCUCAACGCCUGUUACUUCGCAGUGAAGGGGCGUGUGCGUAAAACGCAUAAAA___________ ........................(((....)))..........((((((.(((((....)))))..))))))(((((.....)))))............... (-20.11 = -19.79 + -0.33)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:00:01 2011