
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 27,295,883 – 27,295,981 |
| Length | 98 |
| Max. P | 0.732616 |

| Location | 27,295,883 – 27,295,981 |
|---|---|
| Length | 98 |
| Sequences | 8 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 76.23 |
| Shannon entropy | 0.42714 |
| G+C content | 0.46138 |
| Mean single sequence MFE | -26.20 |
| Consensus MFE | -15.32 |
| Energy contribution | -14.60 |
| Covariance contribution | -0.72 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.732616 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 27295883 98 + 27905053 AAACAAAAGACUGAGGGCACAAACUUGGCCGGAACGCACUUGCUGCGUUUCGCGAGU-------UAUUGUUGGCUCUCGCUUUUCGUUCGUUCAUUUCUUUUGGC ...(((((((..(((((((((((((((..(((((((((.....))))))))))))))-------..))))..))))))((.....)).........))))))).. ( -31.20, z-score = -1.96, R) >droSim1.chr3R 26927690 94 + 27517382 AAACAAAAGACUGAGGGCACAAACUUGGCCGGAACGCACUUGCUGCGUUUCGCGAGU-------UAUUGUUGGCUCUCGCUUUUCGUU----CAUUUCUUUUGGC ...(((((((..(((((((((((((((..(((((((((.....))))))))))))))-------..))))..))))))((.....)).----....))))))).. ( -31.20, z-score = -2.46, R) >droSec1.super_4 6095514 94 + 6179234 AAACAAAAGACUGAGGGCACAAACUUGGCCGGAACGCACUUGCUGCGUUUCGCGAGU-------UAUUGUGGGCUCUCGCUUUUCGUU----CAUUUCUUUUGGC ...(((((((..(((((((((((((((..(((((((((.....))))))))))))))-------..))))..))))))((.....)).----....))))))).. ( -31.20, z-score = -2.24, R) >droYak2.chr3R 28228654 94 + 28832112 AAACAAAAGACUGAGGGCACAAACUUGGCCGGAACGCACUUGCUGCGUUUUGCGAGU-------UAUUGUUGGCUCUCGCUUUUCGUU----CAUUUCUUUUGGC ...(((((((..(((((((((((((((..(((((((((.....))))))))))))))-------..))))..))))))((.....)).----....))))))).. ( -29.20, z-score = -1.89, R) >droEre2.scaffold_4820 9781513 94 - 10470090 AAACAAAAGACUGAGGGCACAAACUUGGCCGGAACGCACUUGCUGCGUUUCGCGAGU-------UAUUGUUGGCUCUCGCUUUUCGUU----CAUUUCUUUUGGC ...(((((((..(((((((((((((((..(((((((((.....))))))))))))))-------..))))..))))))((.....)).----....))))))).. ( -31.20, z-score = -2.46, R) >droAna3.scaffold_13340 16421828 101 + 23697760 AGACAAAAGACUUAGGGCACAAACUUGGCCGGAACGCACUUGAUGCGUUUCGCGAGUGGCGUUUUAUUGUUAGUUCUCGGUUUUUGCC----CAUUUCUUUUGGC ...(((((((..(.(((((.((((.....(((((((((.....)))))))))(((((((((......)))))...)))))))).))))----))..))))))).. ( -36.20, z-score = -3.37, R) >dp4.chr2 4522653 77 + 30794189 AGACAAAAGACUCGAGACACAAACUUGGCCAAAACGCACUUGUUGCGAUUU-CAGCU-------UUUUGCCCAUUUUCUUUGACU-------------------- ...(((((..((.(((((.(((.....((......)).....))).).)))-)))..-------)))))................-------------------- ( -9.70, z-score = 0.90, R) >droPer1.super_19 221367 77 + 1869541 AGACAAAAGACUCGAGACACAAACUUGGCCAAAACGCACUUGUUGCGAUUU-CAGCU-------UUUUGCCCAUUUUCUUUGACU-------------------- ...(((((..((.(((((.(((.....((......)).....))).).)))-)))..-------)))))................-------------------- ( -9.70, z-score = 0.90, R) >consensus AAACAAAAGACUGAGGGCACAAACUUGGCCGGAACGCACUUGCUGCGUUUCGCGAGU_______UAUUGUUGGCUCUCGCUUUUCGUU____CAUUUCUUUUGGC .............((((((((((((((..(((((((((.....)))))))))))))).........))))..)))))............................ (-15.32 = -14.60 + -0.72)
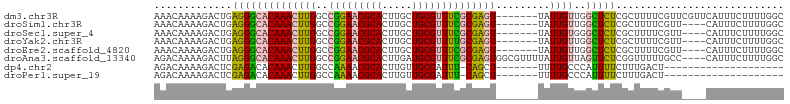
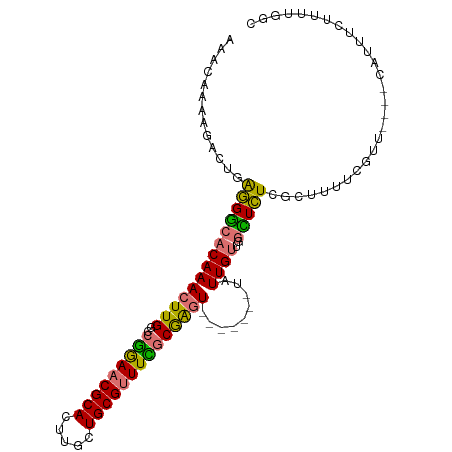

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:59:43 2011