
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 27,189,161 – 27,189,275 |
| Length | 114 |
| Max. P | 0.638799 |

| Location | 27,189,161 – 27,189,275 |
|---|---|
| Length | 114 |
| Sequences | 10 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 84.93 |
| Shannon entropy | 0.31701 |
| G+C content | 0.34702 |
| Mean single sequence MFE | -27.40 |
| Consensus MFE | -15.06 |
| Energy contribution | -15.27 |
| Covariance contribution | 0.21 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.636995 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 27189161 114 + 27905053 -GAGAAACAACAAGCGGG-AGUACAUAGAAACAUUUUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCUAAGCUGGGAAGCAGCUUAGUGAGG-- -(((((((((...(((((-(........(((((((.....)))))))((((......))))..........))))))....)))))))))((((((((.....)))))))).....-- ( -29.60, z-score = -2.90, R) >droAna3.scaffold_13340 5067987 113 + 23697760 -----GGCAGCAGCCAGAGAGUACAUAGAAACAUUUUCGAAAUGUUUGGCAUAAUUUCGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCUAAGCUGGGAAGCAGCUUAGUGAAGUG -----(((....)))......(((...((((((((((((((((...((((........))))........)))))).)))).))))))((((((((((.....)))))))).)).))) ( -25.90, z-score = -0.78, R) >droEre2.scaffold_4820 9675698 114 - 10470090 -GAGAAACAACAAGCGGG-AGUACAUAGAAACAUUUUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCUAAGCCGGGAAGCAGCUUAGUGAGG-- -(((((((((...(((((-(........(((((((.....)))))))((((......))))..........))))))....)))))))))((((((..(....).)))))).....-- ( -25.30, z-score = -1.42, R) >droYak2.chr3R 28119624 116 + 28832112 CAAGAAACAACACGCGGGGAGUACAUAGAAACAUUUUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCUAAGCUGGGAAGCAGCUUAGUGCGG-- ............(((((..((.......(((((((.....)))))))...((((((((((...............))))))))))))..))(((((((.....)))))))..))).-- ( -28.36, z-score = -2.04, R) >droSec1.super_4 5994472 114 + 6179234 -GAGAAACAACAAGCGGG-AGUACAUAGAAACAUUUUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCUAAGCUGGGAAGCAGCUUAGUGAGG-- -(((((((((...(((((-(........(((((((.....)))))))((((......))))..........))))))....)))))))))((((((((.....)))))))).....-- ( -29.60, z-score = -2.90, R) >droMoj3.scaffold_6540 4897421 111 - 34148556 ----UAGCAAGAACAGCC-ACUACAUAGAAACAUUCUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCUCUAAGCUGGAGGCAGCUUAGCAAAG-- ----.((.((((((.(((-(..((((.((.......))...)))).))))..((((((((...............))))))))))))))))((((((((....)))))))).....-- ( -27.46, z-score = -2.21, R) >droGri2.scaffold_14830 5740600 111 - 6267026 ----UAGCAAGAACAGCC-ACUACAUAGAAACAUUCUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCCUAAGCUGGAGGCAGCUUAGUAAAG-- ----....((((((.(((-(..((((.((.......))...)))).))))..((((((((...............))))))))))))))..((((((((....)))))))).....-- ( -27.26, z-score = -2.40, R) >dp4.chr2 4383579 116 + 30794189 CAACAGCCAACAGCCACCGAGUACAUAGAAACAUUUUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCUAAGCUGGGAAGCAGCUUAGUGAAG-- .....(((((((.....((((..............))))...)).)))))((((((((((...............))))))))))..(((((((((((.....)))))))).))).-- ( -29.50, z-score = -2.68, R) >droPer1.super_19 87371 116 + 1869541 CAACAGCCAACAGCCACCGAGUACAUAGAAACAUUUUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCUAAGCUGGGAAGCAGCUUAGUGAAG-- .....(((((((.....((((..............))))...)).)))))((((((((((...............))))))))))..(((((((((((.....)))))))).))).-- ( -29.50, z-score = -2.68, R) >droWil1.scaffold_181108 3226597 101 + 4707319 ---------------CCAGAGCACAUAGAAACAUUCUCCAAAUGUUUGGCAUAAUUUUGGUAUAAAAUAAAUUUUGCAAAAUUGUUUU-CCUAAGCUCGAAAGCAGCUUAGUUGAGC- ---------------.............(((((((.....))))))).(((.(((((...........))))).)))......((((.-.((((((((....).)))))))..))))- ( -21.50, z-score = -1.33, R) >consensus __A_AAACAACAACCGCC_AGUACAUAGAAACAUUUUCGAAAUGUUUGGCAUAAUUUUGCUAUAAAAUAAAUUUUGCAAAAUUGUUUUUCCUAAGCUGGGAAGCAGCUUAGUGAAG__ .....................(((....(((((((.....)))))))(((((((((((((...............)))))))))).(((((.......)))))..)))..)))..... (-15.06 = -15.27 + 0.21)

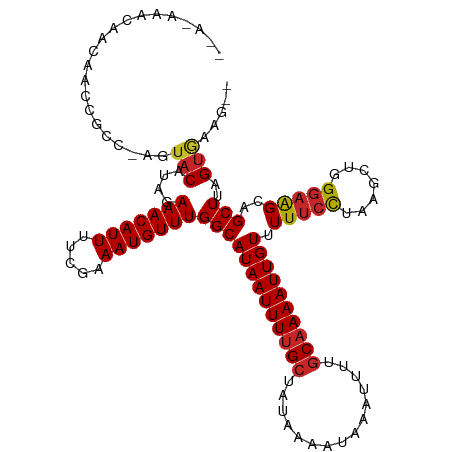
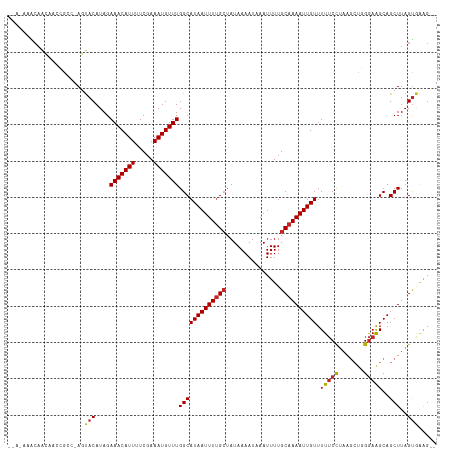
| Location | 27,189,161 – 27,189,275 |
|---|---|
| Length | 114 |
| Sequences | 10 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 84.93 |
| Shannon entropy | 0.31701 |
| G+C content | 0.34702 |
| Mean single sequence MFE | -23.79 |
| Consensus MFE | -14.38 |
| Energy contribution | -14.15 |
| Covariance contribution | -0.23 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.638799 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 27189161 114 - 27905053 --CCUCACUAAGCUGCUUCCCAGCUUAGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAAAAUGUUUCUAUGUACU-CCCGCUUGUUGUUUCUC- --.....((((((((.....))))))))((.(((((((...((...............(((......))).(((((((.....))))))).........-...))..))))))).))- ( -23.30, z-score = -3.07, R) >droAna3.scaffold_13340 5067987 113 - 23697760 CACUUCACUAAGCUGCUUCCCAGCUUAGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCGAAAUUAUGCCAAACAUUUCGAAAAUGUUUCUAUGUACUCUCUGGCUGCUGCC----- ...(((.((((((((.....)))))))))))....((((((((...............)))))))).....(((((((.....))))))).............(((....)))----- ( -23.06, z-score = -1.81, R) >droEre2.scaffold_4820 9675698 114 + 10470090 --CCUCACUAAGCUGCUUCCCGGCUUAGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAAAAUGUUUCUAUGUACU-CCCGCUUGUUGUUUCUC- --.....((((((((.....))))))))((.(((((((...((...............(((......))).(((((((.....))))))).........-...))..))))))).))- ( -22.60, z-score = -2.45, R) >droYak2.chr3R 28119624 116 - 28832112 --CCGCACUAAGCUGCUUCCCAGCUUAGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAAAAUGUUUCUAUGUACUCCCCGCGUGUUGUUUCUUG --.....((((((((.....))))))))((.(((((((..(((...............(((......))).(((((((.....))))))).............))).))))))).)). ( -23.10, z-score = -2.23, R) >droSec1.super_4 5994472 114 - 6179234 --CCUCACUAAGCUGCUUCCCAGCUUAGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAAAAUGUUUCUAUGUACU-CCCGCUUGUUGUUUCUC- --.....((((((((.....))))))))((.(((((((...((...............(((......))).(((((((.....))))))).........-...))..))))))).))- ( -23.30, z-score = -3.07, R) >droMoj3.scaffold_6540 4897421 111 + 34148556 --CUUUGCUAAGCUGCCUCCAGCUUAGAGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAGAAUGUUUCUAUGUAGU-GGCUGUUCUUGCUA---- --.....((((((((....))))))))((((....((((((((...............))))))))..((((.((((...((((...)))).))))..)-)))..)))).....---- ( -24.66, z-score = -1.12, R) >droGri2.scaffold_14830 5740600 111 + 6267026 --CUUUACUAAGCUGCCUCCAGCUUAGGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAGAAUGUUUCUAUGUAGU-GGCUGUUCUUGCUA---- --.....((((((((....))))))))(.((.(((((((((((...............))))))))..((((.((((...((((...)))).))))..)-))).))).)).)..---- ( -25.46, z-score = -1.49, R) >dp4.chr2 4383579 116 - 30794189 --CUUCACUAAGCUGCUUCCCAGCUUAGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAAAAUGUUUCUAUGUACUCGGUGGCUGUUGGCUGUUG --.(((.((((((((.....))))))))))).(((((((((((...............))))))))..(((((.....((((..((((....))))..))))......))))).))). ( -27.86, z-score = -2.14, R) >droPer1.super_19 87371 116 - 1869541 --CUUCACUAAGCUGCUUCCCAGCUUAGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAAAAUGUUUCUAUGUACUCGGUGGCUGUUGGCUGUUG --.(((.((((((((.....))))))))))).(((((((((((...............))))))))..(((((.....((((..((((....))))..))))......))))).))). ( -27.86, z-score = -2.14, R) >droWil1.scaffold_181108 3226597 101 - 4707319 -GCUCAACUAAGCUGCUUUCGAGCUUAGG-AAAACAAUUUUGCAAAAUUUAUUUUAUACCAAAAUUAUGCCAAACAUUUGGAGAAUGUUUCUAUGUGCUCUGG--------------- -((.((.(((((((.......)))))))(-(((...(((((.((((........((((.......)))).......)))).))))).))))..)).)).....--------------- ( -16.66, z-score = 0.10, R) >consensus __CUUCACUAAGCUGCUUCCCAGCUUAGGAAAAACAAUUUUGCAAAAUUUAUUUUAUAGCAAAAUUAUGCCAAACAUUUCGAAAAUGUUUCUAUGUACU_CCCGGUUGUUGUUU_U__ .......(((((((.......))))))).......((((((((...............)))))))).....((((((((...))))))))............................ (-14.38 = -14.15 + -0.23)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:59:29 2011