| Sequence ID | dm3.chr3R |
|---|---|
| Location | 26,885,094 – 26,885,219 |
| Length | 125 |
| Max. P | 0.998498 |

| Location | 26,885,094 – 26,885,219 |
|---|---|
| Length | 125 |
| Sequences | 7 |
| Columns | 134 |
| Reading direction | reverse |
| Mean pairwise identity | 78.69 |
| Shannon entropy | 0.41837 |
| G+C content | 0.40472 |
| Mean single sequence MFE | -29.42 |
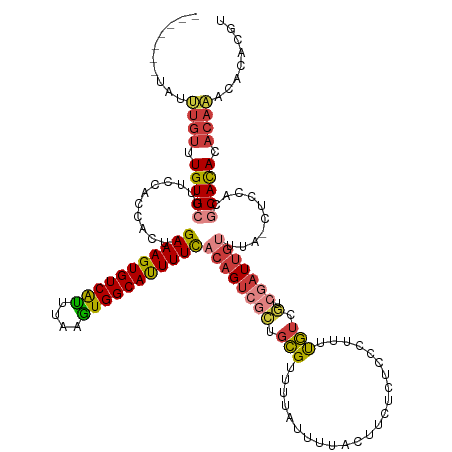
| Consensus MFE | -21.96 |
| Energy contribution | -22.69 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.95 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.38 |
| SVM RNA-class probability | 0.998498 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
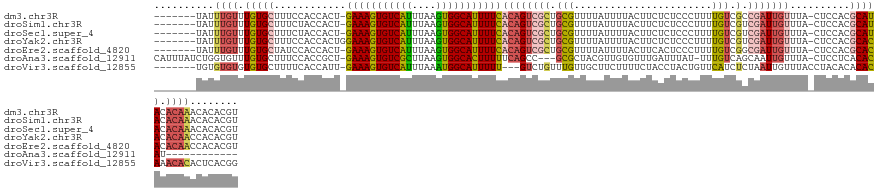
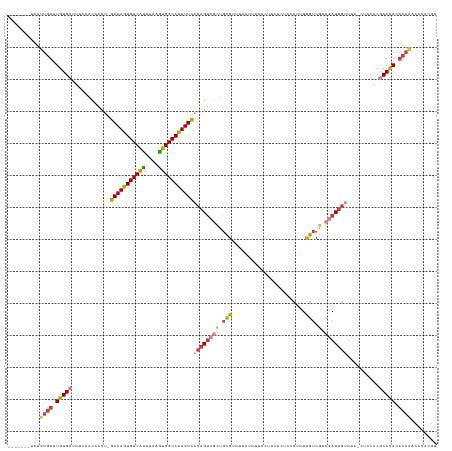
>dm3.chr3R 26885094 125 - 27905053 -------UAUUUGUUUGUGCUUUCCACCACU-GAAAGUGUCAUUUAAGUGGCAUUUUCACAGUCGCUGCGUUUUAUUUUACUUCUCUCCCUUUUGUCGCCGAUUGUUUA-CUCCACGCAUACACAAACACACGU -------..(((((.(((((...........-(((((((((((....)))))))))))(((((((..(((..........................))))))))))...-......))))).)))))....... ( -29.97, z-score = -3.66, R) >droSim1.chr3R 26532559 125 - 27517382 -------UAUUUGUUUGUGCUUUCUACCACU-GAAAGUGUCAUUUAAGUGGCAUUUUCACAGUCGCUGCGUUUUAUUUUACUUCUCUCCCUUUUGUCGUCGAUUGUUUA-CUCCACGCAUACACAAACACACGU -------..(((((.(((((...........-(((((((((((....)))))))))))((((((((.(((.......................))).).)))))))...-......))))).)))))....... ( -29.10, z-score = -3.37, R) >droSec1.super_4 5709641 125 - 6179234 -------UAUUUGUUUGUGCUUUCUACCACU-GAAAGUGUCAUUUAAGUGGCAUUUUCACAGUCGCUGCGUUUUAUUUUACUUCUCUCCCUUUUGUCGUCGAUUGUUUA-CUCCACGCAUACACAAACACACGU -------..(((((.(((((...........-(((((((((((....)))))))))))((((((((.(((.......................))).).)))))))...-......))))).)))))....... ( -29.10, z-score = -3.37, R) >droYak2.chr3R 27824553 126 - 28832112 -------UAUUUGUUUGUGCUUUCCACCACUGGAAAGUGUCAUUUAAGUGGCAUUUUCACAGUCGCUGCGUUUUAUUUUACUUCUCUCCCUUUUGUCGUCGAUUGUUUA-CUCCACGCACACACAACCACACGU -------...((((.(((((............(((((((((((....)))))))))))((((((((.(((.......................))).).)))))))...-......))))).))))........ ( -30.00, z-score = -3.10, R) >droEre2.scaffold_4820 9372773 125 + 10470090 -------UAUUUGUUUGUGCUAUCCACCACU-GAAAGUGUCAUUUAAGUGGCAUUUUCACAGUCGCUGCGUUUUAUUUUACUUCACUCCCUUUUGUCGGCGAUUGUUUA-CUCCACGCACACACAACCACACGU -------...((((.(((((...........-(((((((((((....)))))))))))((((((((((((.......................)).))))))))))...-......))))).))))........ ( -35.50, z-score = -4.55, R) >droAna3.scaffold_12911 1324732 116 + 5364042 CAUUUAUCUGGUGUUUGUGCUUUCCACCGCU-GAAAGUGUCGCUUAAGUGGCACUUUUUCAGCC---GCGCUACGUUGUGUUUGAUUUAU-UUUGUCAGCAAUUGUUUA-CUCCUCACACAU------------ ..........((((..((((........(((-(((((((((((....))))))))..)))))).---)))).(((...((((.(((....-...)))))))..)))...-......))))..------------ ( -27.80, z-score = -1.33, R) >droVir3.scaffold_12855 8701339 123 + 10161210 -------UGUGUGUGUGUGCUUUUCACCAUU-GAAAGUGUCAUUUAAAUGGCAUUUUU---GUCUGUUUGUUGCUUCUUUUCUACCUACUGUUCAUCUCUAAUUGUUUACCUACACACACAAACACACUCACGG -------.(((.((((((((..(..((...(-(((((((((((....)))))))))))---)...))..)..)).............................(((......))).......)))))).))).. ( -24.50, z-score = -1.27, R) >consensus _______UAUUUGUUUGUGCUUUCCACCACU_GAAAGUGUCAUUUAAGUGGCAUUUUCACAGUCGCUGCGUUUUAUUUUACUUCUCUCCCUUUUGUCGUCGAUUGUUUA_CUCCACGCACACACAAACACACGU ..........((((.(((((............(((((((((((....)))))))))))((((((((.(((.......................))).).)))))))..........))))).))))........ (-21.96 = -22.69 + 0.72)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:58:50 2011