
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 26,725,271 – 26,725,403 |
| Length | 132 |
| Max. P | 0.992165 |

| Location | 26,725,271 – 26,725,403 |
|---|---|
| Length | 132 |
| Sequences | 6 |
| Columns | 132 |
| Reading direction | forward |
| Mean pairwise identity | 82.47 |
| Shannon entropy | 0.33863 |
| G+C content | 0.46423 |
| Mean single sequence MFE | -37.45 |
| Consensus MFE | -25.48 |
| Energy contribution | -25.82 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.44 |
| Mean z-score | -2.82 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.52 |
| SVM RNA-class probability | 0.992165 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
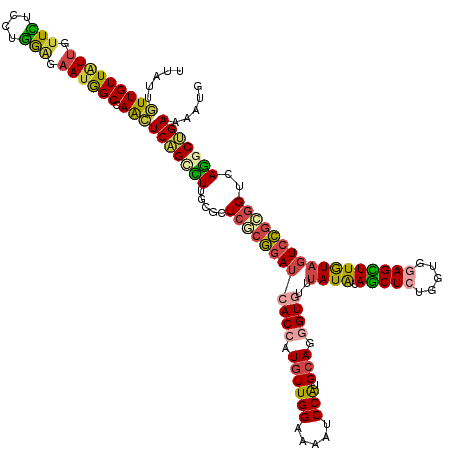
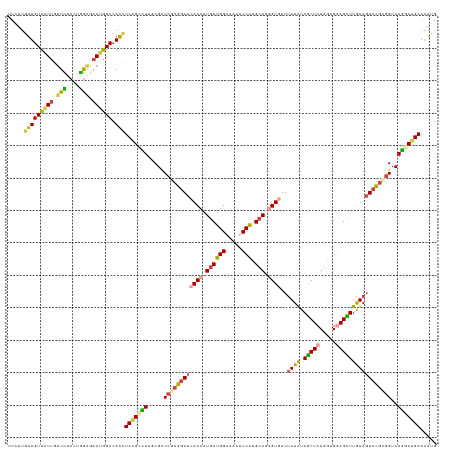
>dm3.chr3R 26725271 132 + 27905053 CAUUUUCAGCCUGACCGCGGAGUACAAGCUCCACCAGAGCUACAUAAACACCCUGCAUGGAUUUUCCAACAUGGUGAUCCGCGAGCGAAAGGCUGAGUUGGCCAUUCUCCAGGAGAACAAUAACAACAAUAA .....(((((((..(((((((.....(((((.....))))).......((((.((..(((.....))).)).)))).)))))).)....)))))))((((....(((((...)))))......))))..... ( -43.90, z-score = -4.26, R) >droWil1.scaffold_181130 10312951 126 + 16660200 CCUCUUCCGCCUGACCGAUGACUAUAAACUGCAUCAGAGCUAUAUCAAUACGCUUCAUGGCUUCUCCAAUAUGGUGAUUAGGGAACGCAAAACGGAGCUGGCUUUGGCCAAUGAUAAUAACAACAA------ ........((((((....((.(........))))))).))..(((((....(((((...(((((((.(((......))).))))).)).....)))))((((....)))).)))))..........------ ( -25.70, z-score = 0.27, R) >droSim1.chr3R 26366854 132 + 27517382 CAUUUUCAGUCUGACCGCGGACUACAAGCUCCACCAGAGCUACAUAAACACCCUGCAUGGAUUUUCCAACAUGGUGAUCCGGGAGCGGAAGGCUGAGUUGGCCAUUCUCCAGGAGAACAAUAACAACAAUAA .....(((((((..((((...((...(((((.....))))).......((((.((..(((.....))).)).))))....))..)))).)))))))((((....(((((...)))))......))))..... ( -39.60, z-score = -2.31, R) >droSec1.super_4 5550928 132 + 6179234 CAUUUUCAGCCUGACCGCGGACUACAAGCUCCACCAGAGCUACAUAAACACCUUGCAUGGAUUUUCCAACAUGGUGAUCCGCGAGCGAAAGGCUGAGUUGGCCAUUCUCCAGGAGAACAAUAACAACAAUAA .....(((((((..(((((((.....(((((.....))))).......((((.((..(((.....))).)).)))).)))))).)....)))))))((((....(((((...)))))......))))..... ( -43.60, z-score = -4.64, R) >droEre2.scaffold_4820 9205267 132 - 10470090 CAUUUUCAGCCUGACCGCGGACUACAAGCUCCACCAGAGCUAUAUAAACACCCUGCAUGGGUUUUCCAACAUGGUGAUCCGCGAACGCAAGGCUGAGCUGGCCAUUCUCCAGGAGAGCAAUAACAACAAUAA .....(((((((...((((((.....(((((.....))))).......((((.((..(((.....))).)).)))).))))))......)))))))(((..((........))..))).............. ( -42.70, z-score = -3.31, R) >droAna3.scaffold_12911 1134986 132 - 5364042 CAUUUUCCGGUUGACUUCGGACUACAAACUUCACCAGAGUUAUAUAAAUACUCUGCACGGAUUUUCCAACAUGGUCAUCCGAGAACGUAAAGCUGAACUCGCCAUUCUUCAGGAAAAUAACAACAAUAACAA .((((((((((.((.(((((..(((.........((((((.........))))))..(((((...((.....))..))))).....)))...))))).)))))........))))))).............. ( -29.20, z-score = -2.68, R) >consensus CAUUUUCAGCCUGACCGCGGACUACAAGCUCCACCAGAGCUACAUAAACACCCUGCAUGGAUUUUCCAACAUGGUGAUCCGCGAACGAAAGGCUGAGCUGGCCAUUCUCCAGGAGAACAAUAACAACAAUAA .....(((((((...((((((.....(((((.....))))).......((((.((..(((.....))).)).)))).))))))......)))))))(((..((........))..))).............. (-25.48 = -25.82 + 0.34)
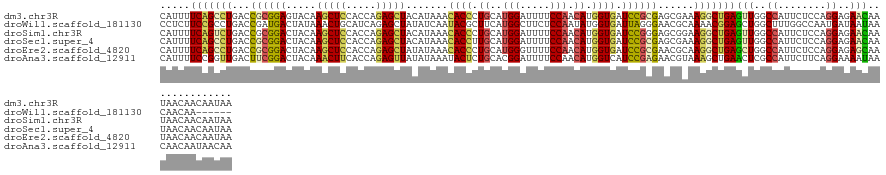
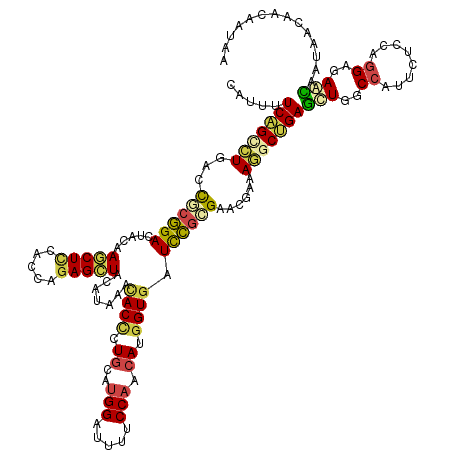
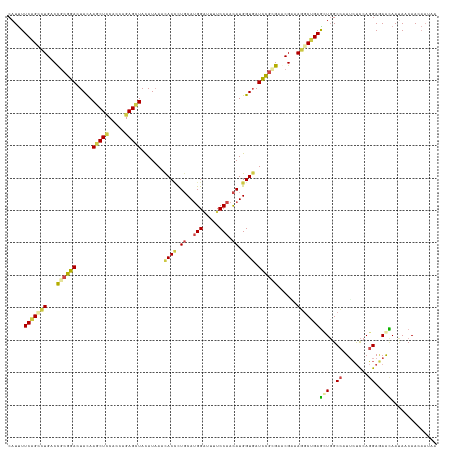
| Location | 26,725,271 – 26,725,403 |
|---|---|
| Length | 132 |
| Sequences | 6 |
| Columns | 132 |
| Reading direction | reverse |
| Mean pairwise identity | 82.47 |
| Shannon entropy | 0.33863 |
| G+C content | 0.46423 |
| Mean single sequence MFE | -41.05 |
| Consensus MFE | -30.85 |
| Energy contribution | -31.00 |
| Covariance contribution | 0.15 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.899458 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 26725271 132 - 27905053 UUAUUGUUGUUAUUGUUCUCCUGGAGAAUGGCCAACUCAGCCUUUCGCUCGCGGAUCACCAUGUUGGAAAAUCCAUGCAGGGUGUUUAUGUAGCUCUGGUGGAGCUUGUACUCCGCGGUCAGGCUGAAAAUG .....(((((((((.(((....))).))))).))))(((((((...((.((((((.((((.((((((.....))).))).))))..((((.(((((.....))))))))).)))))))).)))))))..... ( -48.20, z-score = -3.09, R) >droWil1.scaffold_181130 10312951 126 - 16660200 ------UUGUUGUUAUUAUCAUUGGCCAAAGCCAGCUCCGUUUUGCGUUCCCUAAUCACCAUAUUGGAGAAGCCAUGAAGCGUAUUGAUAUAGCUCUGAUGCAGUUUAUAGUCAUCGGUCAGGCGGAAGAGG ------...............(((((....)))))(((((.....))((((....((.((.....)).)).(((.((((((.(((....))))))(((((((........).))))))))))))))))))). ( -27.00, z-score = 1.18, R) >droSim1.chr3R 26366854 132 - 27517382 UUAUUGUUGUUAUUGUUCUCCUGGAGAAUGGCCAACUCAGCCUUCCGCUCCCGGAUCACCAUGUUGGAAAAUCCAUGCAGGGUGUUUAUGUAGCUCUGGUGGAGCUUGUAGUCCGCGGUCAGACUGAAAAUG .....(((((((((.(((....))).))))).))))((((.((.((((....((((((((.((((((.....))).))).))))..((((.(((((.....)))))))))))))))))..)).))))..... ( -39.40, z-score = -0.84, R) >droSec1.super_4 5550928 132 - 6179234 UUAUUGUUGUUAUUGUUCUCCUGGAGAAUGGCCAACUCAGCCUUUCGCUCGCGGAUCACCAUGUUGGAAAAUCCAUGCAAGGUGUUUAUGUAGCUCUGGUGGAGCUUGUAGUCCGCGGUCAGGCUGAAAAUG .....(((((((((.(((....))).))))).))))(((((((...((.(((((((((((.((((((.....))).))).))))..((((.(((((.....)))))))))))))))))).)))))))..... ( -49.60, z-score = -3.78, R) >droEre2.scaffold_4820 9205267 132 + 10470090 UUAUUGUUGUUAUUGCUCUCCUGGAGAAUGGCCAGCUCAGCCUUGCGUUCGCGGAUCACCAUGUUGGAAAACCCAUGCAGGGUGUUUAUAUAGCUCUGGUGGAGCUUGUAGUCCGCGGUCAGGCUGAAAAUG .....(((((((((.(((.....)))))))).))))(((((((.((...(((((((((((.((((((.....))).))).))))..((((.(((((.....)))))))))))))))))).)))))))..... ( -49.30, z-score = -2.62, R) >droAna3.scaffold_12911 1134986 132 + 5364042 UUGUUAUUGUUGUUAUUUUCCUGAAGAAUGGCGAGUUCAGCUUUACGUUCUCGGAUGACCAUGUUGGAAAAUCCGUGCAGAGUAUUUAUAUAACUCUGGUGAAGUUUGUAGUCCGAAGUCAACCGGAAAAUG .............((((((((.(..((....((..(((((((((((...(.(((((..((.....))...))))).)((((((.........)))))))))))).))).))..))...))..).)))))))) ( -32.80, z-score = -1.34, R) >consensus UUAUUGUUGUUAUUGUUCUCCUGGAGAAUGGCCAACUCAGCCUUGCGCUCGCGGAUCACCAUGUUGGAAAAUCCAUGCAGGGUGUUUAUAUAGCUCUGGUGGAGCUUGUAGUCCGCGGUCAGGCUGAAAAUG .....(((((((((.(((....))).)))))).)))(((((((.....((((((((((((.((((((.....))).))).))))..((((.(((((.....)))))))))))))))))..)))))))..... (-30.85 = -31.00 + 0.15)
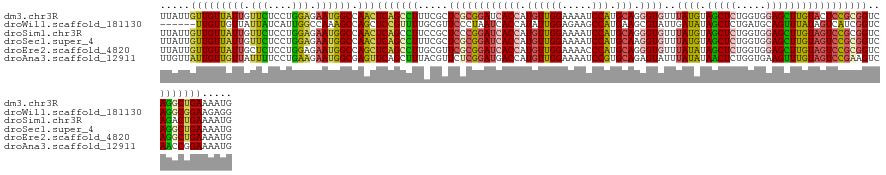
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:58:31 2011