
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 26,686,367 – 26,686,494 |
| Length | 127 |
| Max. P | 0.679331 |

| Location | 26,686,367 – 26,686,494 |
|---|---|
| Length | 127 |
| Sequences | 9 |
| Columns | 129 |
| Reading direction | forward |
| Mean pairwise identity | 69.83 |
| Shannon entropy | 0.63838 |
| G+C content | 0.48231 |
| Mean single sequence MFE | -33.99 |
| Consensus MFE | -16.62 |
| Energy contribution | -16.87 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.30 |
| Mean z-score | -0.84 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.679331 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
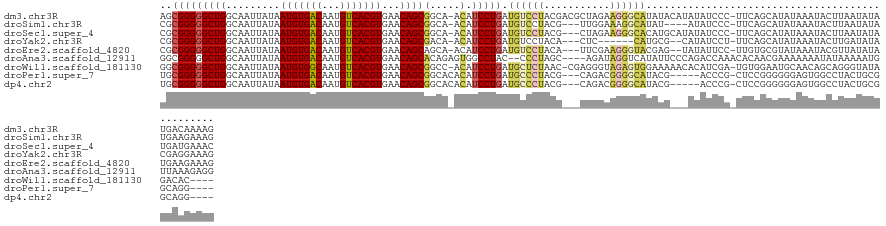
>dm3.chr3R 26686367 127 + 27905053 AGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCGGCA-ACAUCCUGAUGUCCUACGACGCUAGAAGGGCAUAUACAUAUAUCCC-UUCAGCAUAUAAAUACUUAAUAUAUGACAAAAG ..(((((((((.........(((((((...)))))))...))))(...-.).))))).((((.......(((.((((((.((((....)))))))-))))))(((((.........))))))))).... ( -35.50, z-score = -2.52, R) >droSim1.chr3R 26327463 120 + 27517382 CGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCGGCA-ACAUCCUGAUGUCCUACG---UUGGAAAGGCAUAU----AUAUCCC-UUCAGCAUAUAAAUACUUAAUAUAUGAAGAAAG ...((((((((.........(((((((...)))))))...))))(...-.)..(((....(((....---..))).))).....----...))))-(((..((((((.........))))))..))).. ( -27.40, z-score = -0.29, R) >droSec1.super_4 5511756 124 + 6179234 CGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCGGCA-ACAUCCUGAUGUCCUACG---CUAGAAGGGCACAUGCAUAUAUCCC-UUCAGCAUAUAAAUACUUAAUAUAUGAUGAAAC ...((((((((.........(((((((...)))))))...)))).(((-......((.((((((.(.---...).))))))))))).....))))-((((.((((((.........)))))).)))).. ( -31.50, z-score = -0.77, R) >droYak2.chr3R 27609814 116 + 28832112 CGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCGACA-ACAUCCUGAUGUCCUACA---CUC------CAUGCG--CAUAUCCU-UUCAGCAUAUAAAUACUUGAUAUACGAGGAAAG ((((((((.((((..((...(((((((...))))))).))..))((((-.(.....).))))...))---)))------).))))--......((-(((..(.((((.........)))).)..))))) ( -25.50, z-score = 0.02, R) >droEre2.scaffold_4820 9166122 122 - 10470090 CGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCAGCA-ACAUCCUGAUGUCCUACA---UUCGAAGGGUACGAG--UAUAUUCC-UUGUGCGUAUAAAUACGUUAUAUAUGAAGAAAG (((((((((((.(..((...(((((((...))))))).))..))))).-..(((((((((.....))---))...))))).....--......))-)))))((((((.........))))))....... ( -27.60, z-score = 0.30, R) >droAna3.scaffold_12911 1087498 123 - 5364042 GGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCACAGAGUGGCCUAC--CCCUAGC----AGAUAGGUCAUAUUCCCAGACCAAACACAACGAAAAAAAUAUAAAAAUGUUAAAGAGG .(((((((..(((.........(((((...)))))(((.....)))......)))..)--)))).))----.....((((.........)))).................................... ( -30.00, z-score = -1.26, R) >droWil1.scaffold_181130 10265657 122 + 16660200 GGCGGGGGCUGGCAAUUAUAAUGUGGCAAUGUCACGUGAACAGCGGCC-ACAUCCUGAUGCUCUAAC-CGAGGGUAGAGUGGAAAAACACAUCGA-UGUGGAAUGCAACAGCAGGGUAUAGACAC---- .......((((.........(((((((...))))))).....(((.((-(((((....((((((...-..))))))(((((......))).))))-)))))..)))..)))).............---- ( -35.00, z-score = -0.57, R) >droPer1.super_7 1170128 116 - 4445127 UGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCGGCACACAUCCUGAUGCCCUACG---CAGACGGGGCAUACG-----ACCCG-CUCCGGGGGGAGUGGCCUACUGCGGCAGG---- .((((..((((.........(((((((...)))))))...))))(((.(((.((((.(((((((...---.....)))))))...-----.((((-...))))))))))))))..))))......---- ( -46.70, z-score = -1.22, R) >dp4.chr2 30302301 116 - 30794189 UGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCGGCACACAUCCUGAUGCCCUACG---CAGACGGGGCAUACG-----ACCCG-CUCCGGGGGGAGUGGCCUACUGCGGCAGG---- .((((..((((.........(((((((...)))))))...))))(((.(((.((((.(((((((...---.....)))))))...-----.((((-...))))))))))))))..))))......---- ( -46.70, z-score = -1.22, R) >consensus CGCGGGGGCUGGCAAUUAUAAUGUGACAAUGUCACGUGAACAGCGGCA_ACAUCCUGAUGUCCUACG___CUGGAAGGGCAUAUG___AUAUCCC_UUCAGCAUAUAAAUACUUAAUAUAUGAAGAAAG ..(((((((((.........(((((((...)))))))...))))(.....).))))).((((((...........))))))................................................ (-16.62 = -16.87 + 0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:58:23 2011