
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 26,561,742 – 26,561,834 |
| Length | 92 |
| Max. P | 0.869821 |

| Location | 26,561,742 – 26,561,834 |
|---|---|
| Length | 92 |
| Sequences | 11 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 87.25 |
| Shannon entropy | 0.25067 |
| G+C content | 0.41441 |
| Mean single sequence MFE | -24.77 |
| Consensus MFE | -14.92 |
| Energy contribution | -15.02 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.869821 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 26561742 92 + 27905053 -----------CUGGAUGCGAACGACGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG -----------.((..((((((..(((((((.((.......)).)))))............((((((.....))))))..))..)).))))..))........ ( -23.70, z-score = -2.77, R) >droSim1.chr3R 26201196 92 + 27517382 -----------CUGGAUGCGAACGACGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG -----------.((..((((((..(((((((.((.......)).)))))............((((((.....))))))..))..)).))))..))........ ( -23.70, z-score = -2.77, R) >droSec1.super_4 5380358 92 + 6179234 -----------CUGGAUGCGAACGACGCCGCAGUUCAAUAUGCAGCGACAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG -----------.((..((((.....(((.(((........))).)))......((((.((.((((((.....)))))).)).)))).))))..))........ ( -19.70, z-score = -1.81, R) >droYak2.chr3R 27481870 92 + 28832112 -----------CUGGAUGCGAACGACGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG -----------.((..((((((..(((((((.((.......)).)))))............((((((.....))))))..))..)).))))..))........ ( -23.70, z-score = -2.77, R) >droEre2.scaffold_4820 9032550 92 - 10470090 -----------CUGGAUGCGAACGACGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG -----------.((..((((((..(((((((.((.......)).)))))............((((((.....))))))..))..)).))))..))........ ( -23.70, z-score = -2.77, R) >droAna3.scaffold_12911 940681 92 - 5364042 -----------CCAGUGGCAAGCGAUGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG -----------...(((((..((..((((((.((.......)).)))))).))((((.((.((((((.....)))))).)).))))..)).)))......... ( -25.50, z-score = -3.05, R) >dp4.chr2 30148898 97 - 30794189 ------GAGGAGUCGUGCGAUGCGAUGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG ------.....((.(((((.(((..((((((.((.......)).)))))).))).......((((((.....)))))).........)))))))......... ( -28.40, z-score = -2.84, R) >droPer1.super_7 1015265 97 - 4445127 ------GAGGAGUCGUGCGAUGCGAUGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG ------.....((.(((((.(((..((((((.((.......)).)))))).))).......((((((.....)))))).........)))))))......... ( -28.40, z-score = -2.84, R) >droVir3.scaffold_13047 11356907 97 + 19223366 -----GGGAGCGCAGUCAAGAGCU-UGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAA -----....(((.........(((-((((((.((.......)).)))))))))((((.((.((((((.....)))))).)).)))).)))............. ( -27.30, z-score = -3.31, R) >droMoj3.scaffold_6540 3760867 97 - 34148556 -----GGGAGCGCAGUCAAGAGCU-UGCCACAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGUACACAUUACACAG -----..(.(((.........(((-((((.(.((.......)).).)))))))((((.((.((((((.....)))))).)).)))).))).)........... ( -19.70, z-score = -1.22, R) >droGri2.scaffold_14624 1381993 102 + 4233967 AGGGAGGGAGCGCUGUGAAGAGCU-UGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAC .........(((..((((...(((-((((((.((.......)).)))))))))........((((((.....))))))...))))..)))............. ( -28.70, z-score = -2.58, R) >consensus ___________CUGGUUGCGAGCGAUGCCGCAGUUCAAUAUGCAGCGGCAGGCAAUAUUAAAAUAUCUUAAGGAUAUUAUGUUAUUACGCACACAUUACACAG ..........................(((((.((.......)).)))))..((((((.((.((((((.....)))))).)).))))..))............. (-14.92 = -15.02 + 0.10)


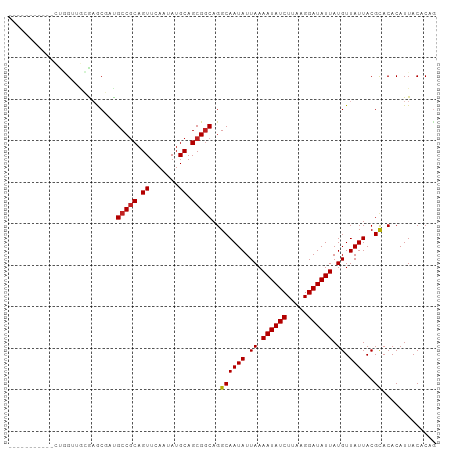
| Location | 26,561,742 – 26,561,834 |
|---|---|
| Length | 92 |
| Sequences | 11 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 87.25 |
| Shannon entropy | 0.25067 |
| G+C content | 0.41441 |
| Mean single sequence MFE | -24.68 |
| Consensus MFE | -17.30 |
| Energy contribution | -16.77 |
| Covariance contribution | -0.53 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.841384 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 26561742 92 - 27905053 CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCGUCGUUCGCAUCCAG----------- (((.(..(((((.((((((((....((((((.....))))))....))))))..(.(((((((........))))))).))).)))))))))----------- ( -24.20, z-score = -1.96, R) >droSim1.chr3R 26201196 92 - 27517382 CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCGUCGUUCGCAUCCAG----------- (((.(..(((((.((((((((....((((((.....))))))....))))))..(.(((((((........))))))).))).)))))))))----------- ( -24.20, z-score = -1.96, R) >droSec1.super_4 5380358 92 - 6179234 CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGUCGCUGCAUAUUGAACUGCGGCGUCGUUCGCAUCCAG----------- (((.(..(((((.((((((((....((((((.....))))))....))))))..(.(((((((........))))))).))).)))))))))----------- ( -25.30, z-score = -2.54, R) >droYak2.chr3R 27481870 92 - 28832112 CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCGUCGUUCGCAUCCAG----------- (((.(..(((((.((((((((....((((((.....))))))....))))))..(.(((((((........))))))).))).)))))))))----------- ( -24.20, z-score = -1.96, R) >droEre2.scaffold_4820 9032550 92 + 10470090 CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCGUCGUUCGCAUCCAG----------- (((.(..(((((.((((((((....((((((.....))))))....))))))..(.(((((((........))))))).))).)))))))))----------- ( -24.20, z-score = -1.96, R) >droAna3.scaffold_12911 940681 92 + 5364042 CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCAUCGCUUGCCACUGG----------- .........(((.((((((((....((((((.....))))))....)))))).((((((.............))))))......)))))...----------- ( -22.62, z-score = -1.21, R) >dp4.chr2 30148898 97 + 30794189 CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCAUCGCAUCGCACGACUCCUC------ .((((..(((((...((((((....((((((.....))))))....)))))).((((((.............)))))).)))))..)))).......------ ( -27.92, z-score = -3.14, R) >droPer1.super_7 1015265 97 + 4445127 CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCAUCGCAUCGCACGACUCCUC------ .((((..(((((...((((((....((((((.....))))))....)))))).((((((.............)))))).)))))..)))).......------ ( -27.92, z-score = -3.14, R) >droVir3.scaffold_13047 11356907 97 - 19223366 UUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCA-AGCUCUUGACUGCGCUCCC----- ..(((((....(.((((((((....((((((.....))))))....)))))).((((((.............))))))-.)).).....)))))....----- ( -24.02, z-score = -1.99, R) >droMoj3.scaffold_6540 3760867 97 + 34148556 CUGUGUAAUGUGUACGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGUGGCA-AGCUCUUGACUGCGCUCCC----- ..(((((..(.(((.((((((....((((((.....))))))....)))))).))))((..((........))..)).-..........)))))....----- ( -20.60, z-score = -1.48, R) >droGri2.scaffold_14624 1381993 102 - 4233967 GUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCA-AGCUCUUCACAGCGCUCCCUCCCU ..((((..((((.((((((((....((((((.....))))))....)))))).((((((.............))))))-.))....))))))))......... ( -26.32, z-score = -2.57, R) >consensus CUGUGUAAUGUGUGCGUAAUAACAUAAUAUCCUUAAGAUAUUUUAAUAUUGCCUGCCGCUGCAUAUUGAACUGCGGCAUCGCUCGCAACCAG___________ .............((((((((....((((((.....))))))....)))))).((((((.............))))))..))..................... (-17.30 = -16.77 + -0.53)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:58:01 2011