| Sequence ID | dm3.chr3R |
|---|---|
| Location | 26,478,361 – 26,478,464 |
| Length | 103 |
| Max. P | 0.979932 |

| Location | 26,478,361 – 26,478,464 |
|---|---|
| Length | 103 |
| Sequences | 9 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 81.74 |
| Shannon entropy | 0.38699 |
| G+C content | 0.45475 |
| Mean single sequence MFE | -18.74 |
| Consensus MFE | -11.39 |
| Energy contribution | -11.17 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.03 |
| SVM RNA-class probability | 0.979932 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
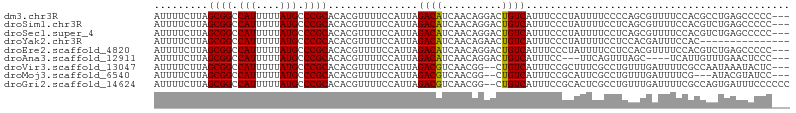
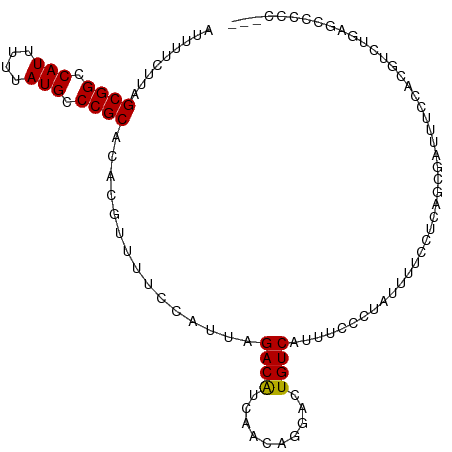
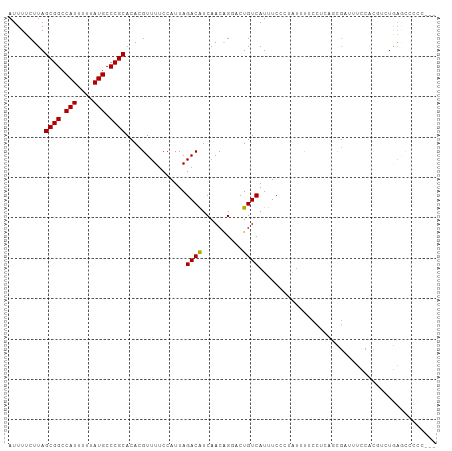
>dm3.chr3R 26478361 103 + 27905053 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACAUCAACAGGACUGUCAUUUCCCUAUUUUCCCCAGCGUUUUCCACGCCUGAGCCCCC--- .........((((.(((....))).))))...............((((((.....)).))))................((((((.....))).))).......--- ( -17.40, z-score = -2.20, R) >droSim1.chr3R 26124272 103 + 27517382 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACAUCAACAGGACUGUCAUUUCCCUAUUUUCCUCAGCGUUUUCCACGUCUGAGCCCCC--- .........((((.(((....))).))))...............((((((.....)).))))..............((((((((.....))).))))).....--- ( -21.20, z-score = -3.66, R) >droSec1.super_4 5304878 103 + 6179234 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACAUCAACAGGACUGUCAUUUCCCUAUUUUCCUCAGCGUUUUCCACGUCUGAGCCCCC--- .........((((.(((....))).))))...............((((((.....)).))))..............((((((((.....))).))))).....--- ( -21.20, z-score = -3.66, R) >droYak2.chr3R 27402477 91 + 28832112 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACAUCAACAGAACUGUCAUUUCCCUAUUUUCCUCCACGAUUUCCAC--------------- .........((((.(((....))).))))...............((((((....))..)))).............................--------------- ( -12.10, z-score = -2.61, R) >droEre2.scaffold_4820 8951489 103 - 10470090 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACAUCAACAGGACUGUCAUUUCCCUAUUUUCCUCCACGUUUUCCACGUCUGAGCCCCC--- .........((((.(((....))).))))...............((((((.....)).))))..............(((.((((.....))))..))).....--- ( -16.60, z-score = -2.27, R) >droAna3.scaffold_12911 859016 96 - 5364042 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACAUCAACAGGACUGUCAUUUCC---UUCAGUUUAGC----UCAUUGUUUGAACUCCC--- .........((((.(((....))).))))............(((((((....(..(((((........---..)))))..).----....)))))))......--- ( -17.70, z-score = -2.23, R) >droVir3.scaffold_13047 11266760 101 + 19223366 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACGUCAACGG--CUGUCAUUUCCGCUUUCGCCUGUUUGAUUUUCGCCAAUAAAUACUC--- .........((((.(((....))).))))..((((((......))))))....((--(.((((..(..((....))..)..))))....)))...........--- ( -21.00, z-score = -2.16, R) >droMoj3.scaffold_6540 3664285 98 - 34148556 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACGUCAACGG--CUGUCAUUUCCGCAUUCGCCUGUUUGAUUUUCG---AUACGUAUCC--- .........((((.(((....))).))))((((((((......))))))...(((--..((((..(..((....))..)..))))..)))---....))....--- ( -19.70, z-score = -1.40, R) >droGri2.scaffold_14624 1255541 104 + 4233967 AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACGUCAACGG--CUGUCAUUUCCGCACUCGCCUGUUUGAUUUUCGCCAGUGAUUUCCCCCC .........((((.(((....))).)))).(((((((......))))......((--(.((((..(..((....))..)..))))....))).))).......... ( -21.80, z-score = -2.15, R) >consensus AUUUUCUUAGCGGCCAUUUUUAUGCCCGCACACGUUUUCCAUUAGACAUCAACAGGACUGUCAUUUCCCUAUUUUCCUCAGCGAUUUCCACGUCUGAGCCCCC___ .........((((.(((....))).))))...............((((..........))))............................................ (-11.39 = -11.17 + -0.22)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:57:46 2011