
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 26,387,659 – 26,387,751 |
| Length | 92 |
| Max. P | 0.664490 |

| Location | 26,387,659 – 26,387,751 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 95.65 |
| Shannon entropy | 0.05989 |
| G+C content | 0.48913 |
| Mean single sequence MFE | -30.83 |
| Consensus MFE | -28.03 |
| Energy contribution | -29.03 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.664490 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
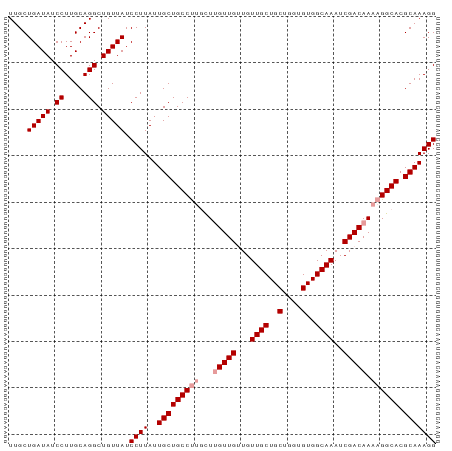
>dm3.chr3R 26387659 92 - 27905053 UUGCUGAUAUCCUUGCAGGCUGUUAUCCUUAUUGCUGCCCUGUUUGUUGCUGUUGCUGCUGGUGUGGCAAAUCGACAAAAGGCACGCAAAGG ....(((((.((.....)).))))).((((..(((((((...(((((((...((((..(....)..))))..))))))).)))).))))))) ( -30.20, z-score = -1.42, R) >droSec1.super_4 5214464 92 - 6179234 UUGCUGAUAUCCUUGCAGGCUGUUAUCCUUAUUGCUGCCUUGCUGGUUGUUGUUGCUGCUGGUGUGGCAAAUCGACAAAAGGCACGCAAAGG ....(((((.((.....)).))))).((((..(((((((((....((((...((((..(....)..))))..))))..)))))).))))))) ( -30.80, z-score = -1.40, R) >droSim1.chr3R 26033080 92 - 27517382 UUGCUGAUAUCCUUGCAGGCUGUUAUCCUUAUUGCUGCCUGCCUUGUUGUUGUUGCUGCUGGUGUGGCAAAUCGACAAAAGGCACGCAAAGG ((((((....((((((((((.((..........)).)))))).((((((...((((..(....)..))))..)))))))))))).))))... ( -31.50, z-score = -1.87, R) >consensus UUGCUGAUAUCCUUGCAGGCUGUUAUCCUUAUUGCUGCCUUGCUUGUUGUUGUUGCUGCUGGUGUGGCAAAUCGACAAAAGGCACGCAAAGG ....(((((.((.....)).))))).((((..(((((((((...(((((...((((..(....)..))))..))))).)))))).))))))) (-28.03 = -29.03 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:57:34 2011