
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 26,300,912 – 26,301,002 |
| Length | 90 |
| Max. P | 0.752114 |

| Location | 26,300,912 – 26,301,002 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 69.48 |
| Shannon entropy | 0.62558 |
| G+C content | 0.50993 |
| Mean single sequence MFE | -24.67 |
| Consensus MFE | -13.18 |
| Energy contribution | -13.04 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.45 |
| Mean z-score | -0.81 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.752114 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
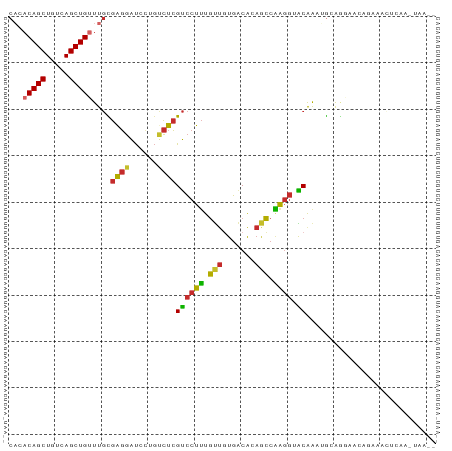
>dm3.chr3R 26300912 90 - 27905053 CACACAGCUGUCAGCUGUUUGCGAGGAUCCUGUCUCGUCCUUUGUUGUGACACAACCAAGGUACAAAUGCAGGAACAGAAACUCAACUAA-- ((.(((((.....))))).)).(((..((((((...((((((.((((.....)))).)))).))....)))))).......)))......-- ( -25.20, z-score = -1.09, R) >droSim1.chr3R 25946734 90 - 27517382 CACACAGCUGUCAGCUGUUUGCGAGGAUCCUGUCUCGUCCUUUGUUGUGACACAGCCAAGGUACAAAUGCAGGAACAGAAACUCAACUAA-- ((.(((((.....))))).)).(((..((((((...((((((.((((.....)))).)))).))....)))))).......)))......-- ( -25.20, z-score = -0.73, R) >droSec1.super_4 5125184 90 - 6179234 CACACAGCUGUCAGCUGUUUGCGAGGAUCCUGUCUCGUCCUUUGUUGUGACACAGCCAAGGUACAAAUGCAGGAACAGAAACUCAAGUAA-- ((.(((((.....))))).)).(((..((((((...((((((.((((.....)))).)))).))....)))))).......)))......-- ( -25.20, z-score = -0.61, R) >droYak2.chr3R 27219794 90 - 28832112 CACACAGCUGUCAGCUGUUUGCGAGGAUCCUGUCUCGUCCUUUGUUGUGACACAGCCAACGUACAAAUGCAGCAACAGAAACUGAAGUAA-- ....(((((((..(((((.(((((((((........)))))..((((.....))))...)))).....))))).))))...)))......-- ( -20.40, z-score = 0.83, R) >droEre2.scaffold_4820 8761215 90 + 10470090 CACACAGCUGUCAGCUGUUUGCGAGGAUCCUGUCUCGUCCUUUGUUGUGCCACAGCCAAGGCACGAAUGCAGAAACAGAAACCGAAGUAA-- ...(((((.....)))))((((..((...((((.((........(((((((........))))))).....)).))))...))...))))-- ( -25.02, z-score = -0.45, R) >droAna3.scaffold_12911 665357 92 + 5364042 CUCACAGCUGUCAGCUGUUUGUGAGGAUCCUGUCUCGUCCUUUGUUAUGACAAAGCCAGGGGACAGCAACAGGAACACAGAUACAGAUACAA ....((((.....))))((((((....((((((((.((((((((............))))))))))..)))))).))))))........... ( -26.10, z-score = -0.73, R) >dp4.chr2 29568459 70 + 30794189 CAAACAGCUGUCAGCUG-UUGUGAGGAUCUCGCCUCGUCCUUUGUUGUGACAAAGCCGAGGAGCGACAGGA--------------------- .......(((((.(((.-....(((...))).(((((..((((((....)))))).)))))))))))))..--------------------- ( -23.60, z-score = -1.07, R) >droPer1.super_7 418711 70 + 4445127 CAAACAGCUGUCAGCUG-UUGUGAGGAUCUCGCCUCGUCCUUUGUUGUGACAAAGCCGAGGAGCGACAGGA--------------------- .......(((((.(((.-....(((...))).(((((..((((((....)))))).)))))))))))))..--------------------- ( -23.60, z-score = -1.07, R) >droMoj3.scaffold_6540 19373083 70 - 34148556 ---ACAGCUGACAGCUGUUU-UGGCAGCCCCCCUCGGCGCUGUCCCUUUGCGUUUGCCAGACGCAAAGACGAGG------------------ ---(((((.....)))))..-.((((((((.....)).)))))).(((((((((.....)))))))))......------------------ ( -27.70, z-score = -2.38, R) >consensus CACACAGCUGUCAGCUGUUUGCGAGGAUCCUGUCUCGUCCUUUGUUGUGACACAGCCAAGGUACAAAUGCAGGAACAGAAACUCAA_UAA__ ((.(((((.....))))).)).((((......))))((((((.(((.......))).)))).))............................ (-13.18 = -13.04 + -0.14)

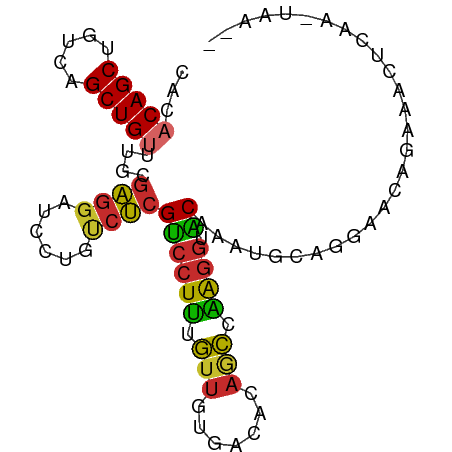
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:57:21 2011