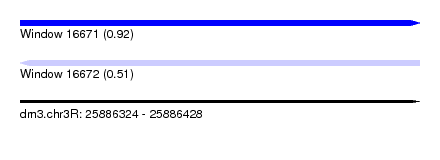

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 25,886,324 – 25,886,428 |
| Length | 104 |
| Max. P | 0.916992 |

| Location | 25,886,324 – 25,886,428 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 78.90 |
| Shannon entropy | 0.37304 |
| G+C content | 0.41842 |
| Mean single sequence MFE | -22.42 |
| Consensus MFE | -17.00 |
| Energy contribution | -18.08 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.916992 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
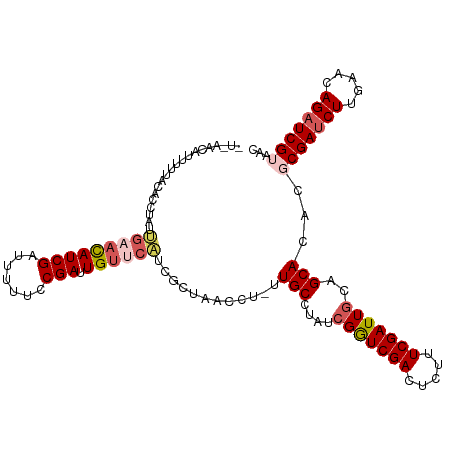
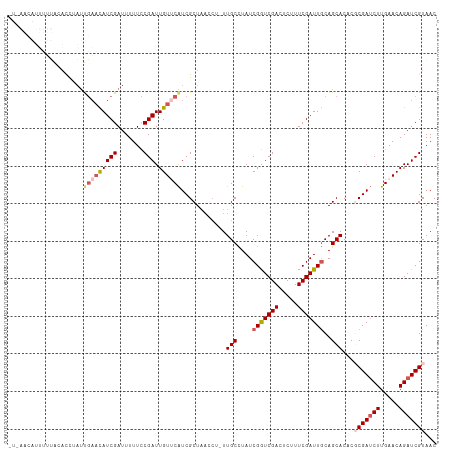
>dm3.chr3R 25886324 104 + 27905053 ----------CCAAACAUUUUGAACAUCGAUUUUUCCGAUUGUUCAUCGCUAACCUUUUGCCUAUCGGUCGACUCUUUCGAUUGCAGCACACGCGAUCUUGAACAGAUCGUAAC ----------..........(((((((((.......))).))))))............(((....(((((((.....)))))))..)))...(((((((.....)))))))... ( -23.40, z-score = -1.93, R) >droSim1.chr3R 25552926 108 + 27517382 CUAAACAUUUUUACACCAAUUGAACAUCGAUUUUUCCGAUUGUUCAUCGCU------UUGCCUAUCGGUCGACUCUUUCGAUUGCAGCACACGCGAUCUUGAACAGAUCGUAAC ..........((((...(((((.....))))).....(((((((((((((.------.(((....(((((((.....)))))))..)))...))))...)))))).))))))). ( -24.00, z-score = -2.15, R) >droSec1.super_4 4732350 113 + 6179234 AUAAACAUUUUUACACCUAUUGAACAUCGAUUUUUCCGAUUGUUCAUCGCUAACCU-UUGCCUAUCGGUCGACUCUUUCGAUUGCAGCACGCGCGAUCUUGAACAGAUCGUAAC ....................(((((((((.......))).))))))..........-.(((....(((((((.....)))))))..)))...(((((((.....)))))))... ( -23.40, z-score = -1.63, R) >droYak2.chr3R 26809507 98 + 28832112 ---------------CUGAACACUUAUCGAUUUUUUCGAUUGUCCGUCGCUAAACUUUUGCUUAUCGAUCGACGCUUUCGAUUGCAGCACUC-CGAACUUGAACAGAUCGUAAC ---------------............((((((.(((((..(((.((((.(((.(....).))).)))).)))((........)).......-.....))))).)))))).... ( -18.00, z-score = -0.05, R) >droEre2.scaffold_4820 8356641 113 - 10470090 CUUAACAUUUUUGUAUCUAUCGAAUAUCGAUUUUUUCGAUUGUCCAUCGCUAACCCCUUGCUUAUCGAUCGACGCUUUCGAUUACAGCACUC-CGAUCUUGAACAGAUCGUAAC ..........(((((...((((((.(((((.....))))).(((.((((.(((........))).)))).)))...))))))))))).....-((((((.....)))))).... ( -23.30, z-score = -2.37, R) >consensus _U_AACAUUUUUACACCUAUUGAACAUCGAUUUUUCCGAUUGUUCAUCGCUAACCU_UUGCCUAUCGGUCGACUCUUUCGAUUGCAGCACACGCGAUCUUGAACAGAUCGUAAC ....................(((((((((.......))).))))))............(((....(((((((.....)))))))..)))...(((((((.....)))))))... (-17.00 = -18.08 + 1.08)

| Location | 25,886,324 – 25,886,428 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 78.90 |
| Shannon entropy | 0.37304 |
| G+C content | 0.41842 |
| Mean single sequence MFE | -26.74 |
| Consensus MFE | -20.26 |
| Energy contribution | -20.14 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.24 |
| Mean z-score | -0.97 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.514449 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
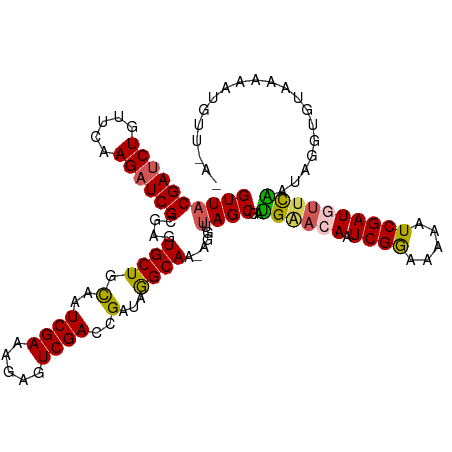
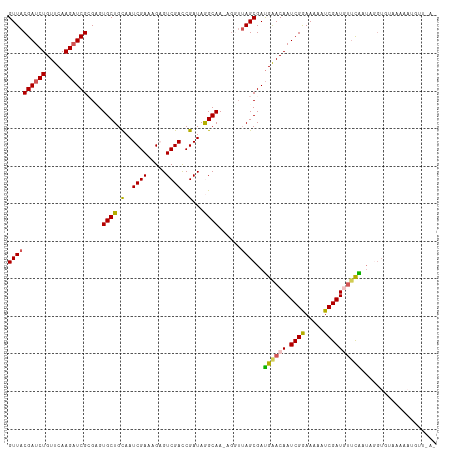
>dm3.chr3R 25886324 104 - 27905053 GUUACGAUCUGUUCAAGAUCGCGUGUGCUGCAAUCGAAAGAGUCGACCGAUAGGCAAAAGGUUAGCGAUGAACAAUCGGAAAAAUCGAUGUUCAAAAUGUUUGG---------- ((((((((((.....))))))(...((((....((....))(((....))).))))...)..))))..((((((.((((.....))))))))))..........---------- ( -24.70, z-score = -0.19, R) >droSim1.chr3R 25552926 108 - 27517382 GUUACGAUCUGUUCAAGAUCGCGUGUGCUGCAAUCGAAAGAGUCGACCGAUAGGCAA------AGCGAUGAACAAUCGGAAAAAUCGAUGUUCAAUUGGUGUAAAAAUGUUUAG ....((((.((((((...((((...((((....((....))(((....))).)))).------.)))))))))))))).....((((((.....)))))).............. ( -27.50, z-score = -1.09, R) >droSec1.super_4 4732350 113 - 6179234 GUUACGAUCUGUUCAAGAUCGCGCGUGCUGCAAUCGAAAGAGUCGACCGAUAGGCAA-AGGUUAGCGAUGAACAAUCGGAAAAAUCGAUGUUCAAUAGGUGUAAAAAUGUUUAU .((((.(((((((....(((((((.(..(((..((....))(((....)))..))).-).))..)))))(((((.((((.....)))))))))))))))))))).......... ( -27.80, z-score = -0.78, R) >droYak2.chr3R 26809507 98 - 28832112 GUUACGAUCUGUUCAAGUUCG-GAGUGCUGCAAUCGAAAGCGUCGAUCGAUAAGCAAAAGUUUAGCGACGGACAAUCGAAAAAAUCGAUAAGUGUUCAG--------------- ....(((.((.....)).)))-(((..((((........))(((..(((.(((((....))))).)))..))).(((((.....))))).))..)))..--------------- ( -23.80, z-score = -0.52, R) >droEre2.scaffold_4820 8356641 113 + 10470090 GUUACGAUCUGUUCAAGAUCG-GAGUGCUGUAAUCGAAAGCGUCGAUCGAUAAGCAAGGGGUUAGCGAUGGACAAUCGAAAAAAUCGAUAUUCGAUAGAUACAAAAAUGUUAAG ....((((((.....))))))-......((((((((((...(((.((((.(((.(....).))).)))).))).(((((.....))))).))))))...))))........... ( -29.90, z-score = -2.25, R) >consensus GUUACGAUCUGUUCAAGAUCGCGAGUGCUGCAAUCGAAAGAGUCGACCGAUAGGCAA_AGGUUAGCGAUGAACAAUCGGAAAAAUCGAUGUUCAAUAGGUGUAAAAAUGUU_A_ ((((((((((.....))))))....((((.(..((((.....))))..)...))))......))))..((((((.((((.....)))))))))).................... (-20.26 = -20.14 + -0.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:56:14 2011