
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 9,170,093 – 9,170,188 |
| Length | 95 |
| Max. P | 0.570964 |

| Location | 9,170,093 – 9,170,188 |
|---|---|
| Length | 95 |
| Sequences | 10 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 76.73 |
| Shannon entropy | 0.47870 |
| G+C content | 0.39256 |
| Mean single sequence MFE | -20.23 |
| Consensus MFE | -7.72 |
| Energy contribution | -7.63 |
| Covariance contribution | -0.09 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.570964 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 9170093 95 + 23011544 UUGUUCCG----CGACCAGAUGG--CUGGCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC--AAAAAGCCAGUU--CAUCUACAC-- ........----.....((((((--(((((....(((.....)))....(((((((...(((......)))..)))))))--.....))))).)--)))))....-- ( -23.20, z-score = -3.09, R) >droSim1.chr2L 8950980 95 + 22036055 UUGUUCCG----CGUCCAGAUG---CUGGCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC--AAAAAGCCAGUUU-CAUCUACAC-- ........----......((.(---(((((....(((.....)))....(((((((...(((......)))..)))))))--.....)))))).)-)........-- ( -19.90, z-score = -2.32, R) >droSec1.super_3 4635546 95 + 7220098 UUGUUCCG----CAUCCAGAUG---CUGGCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC--AAAAAGCCAGUUU-CAUCUACAC-- ........----......((.(---(((((....(((.....)))....(((((((...(((......)))..)))))))--.....)))))).)-)........-- ( -19.90, z-score = -2.70, R) >droYak2.chr2L 11838659 104 + 22324452 CUGUUCCCGAGAUGCCCAGAUGAUGCUGGCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC--AAAAAGCCAGUUU-CAUCUACACAC .................((((((.((((((....(((.....)))....(((((((...(((......)))..)))))))--.....)))))).)-)))))...... ( -26.20, z-score = -3.30, R) >droEre2.scaffold_4929 9775694 97 + 26641161 UUGUUCCG----AGUCUGGAUG---CUGGCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC--AAAAAGCCAGUUU-CAUCUACACGA ........----.((.((((((---(((((....(((.....)))....(((((((...(((......)))..)))))))--.....)))))...-)))))).)).. ( -22.70, z-score = -2.25, R) >dp4.chr4_group5 1480238 96 + 2436548 ---UCUCU----CUCUCUGUUGA--GUCGCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGUGGAAAAAGCCAGUUUUCAUCUACAC-- ---.....----..((..(((((--(............))))))..))((((((((...(((......)))..))))))))(((((......)))))........-- ( -14.00, z-score = -0.60, R) >droWil1.scaffold_180708 133742 78 + 12563649 --------------------UCGCUCAUUCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC------AGCCAGGCU-AACAUACAU-- --------------------...........................(((((((((...(((......)))..)))))))------.))......-.........-- ( -10.40, z-score = -1.50, R) >droVir3.scaffold_12963 550777 95 + 20206255 AAGUAGCCG---CGUCAUGUGCGGCCUUUCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC------AGCGAACUU-CAUCUUG-GC- ((((.((((---((.....))))))......................(((((((((...(((......)))..)))))))------.))..))))-.......-..- ( -21.70, z-score = -1.36, R) >droMoj3.scaffold_6500 12055758 96 + 32352404 AAGUAGCCG---CGUCAUGUGCGGCCUUUCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC------AGCGAACUU-CAUCUUUUGC- ((((.((((---((.....))))))......................(((((((((...(((......)))..)))))))------.))..))))-..........- ( -21.70, z-score = -1.60, R) >droGri2.scaffold_15126 5329932 95 - 8399593 AAGUAGCCG---CGUCAUGUGCGGCCUUUCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC------AGCGAAGUU-CAUCUUG-GC- .....((((---((.....))))))........................(((((((...(((......)))..)))))))------.((.(((..-...))).-))- ( -22.60, z-score = -1.49, R) >consensus UUGUUCCC____CGUCCAGAUGG__CUGGCAAACAAUUUUCAAUUAAGCGCACAAUCACGAUAAAUAAAUCUCAUUGUGC__AAAAAGCCAGUUU_CAUCUACAC__ .................................................(((((((...(((......)))..)))))))........................... ( -7.72 = -7.63 + -0.09)

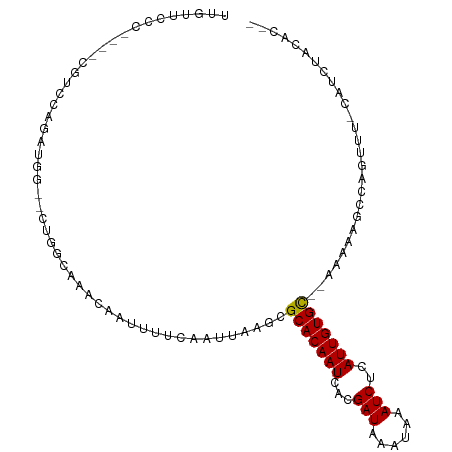

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:27:41 2011