
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 25,798,243 – 25,798,298 |
| Length | 55 |
| Max. P | 0.500000 |

| Location | 25,798,243 – 25,798,298 |
|---|---|
| Length | 55 |
| Sequences | 4 |
| Columns | 55 |
| Reading direction | reverse |
| Mean pairwise identity | 88.48 |
| Shannon entropy | 0.18953 |
| G+C content | 0.42273 |
| Mean single sequence MFE | -11.53 |
| Consensus MFE | -9.46 |
| Energy contribution | -9.15 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
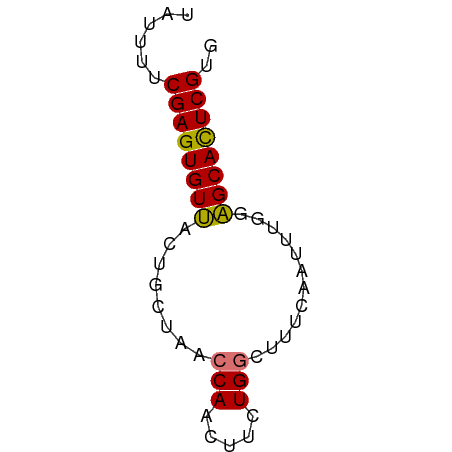
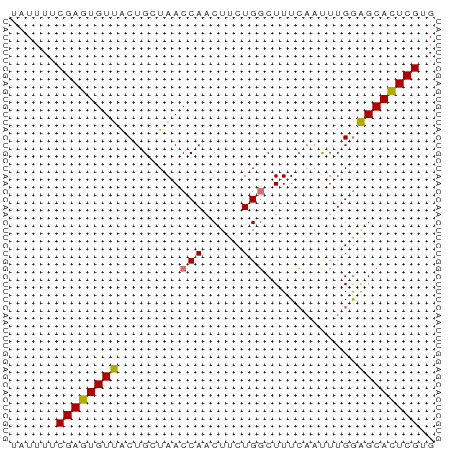
>dm3.chr3R 25798243 55 - 27905053 UAUUUUCGAGUGUUACUGCUAACCAACUUCUGGCUUUCAUCUUGGAGCAUUCGUG ......((((((((.((((((.........)))).........)))))))))).. ( -8.70, z-score = -0.39, R) >droYak2.chr3R 8155050 55 + 28832112 UAUUUUCGAGUGUCGCUGGUAACCAACUUCUGGCUUUCCAUUUGAGGCACUCGUG ......((((((((..(((.(((((.....))).)).))).....)))))))).. ( -16.50, z-score = -2.68, R) >droSec1.super_4 4643764 55 - 6179234 UAUUUUCGAGUGUUACUGCUAACCAACUUCUGUCUUUAAAUUUGGAGCACUCGUG ......((((((((...........((....))............)))))))).. ( -9.40, z-score = -1.17, R) >droSim1.chr3R_random 1247017 55 - 1307089 UAUUUUCGAGUGUUACUGUUAACCACCUUCUGGCUUUCAAUUUGGAGCACUCGUG ......((((((((...(((.....((....)).....)))....)))))))).. ( -11.50, z-score = -1.57, R) >consensus UAUUUUCGAGUGUUACUGCUAACCAACUUCUGGCUUUCAAUUUGGAGCACUCGUG ......((((((((........(((.....)))............)))))))).. ( -9.46 = -9.15 + -0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:56:02 2011